Abstract
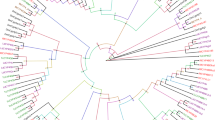
NADPH–cytochrome P450 reductase (CPR) is the most important redox partner of various cytochrome P450 monooxygenases (P450s) and plays a central role in multiple metabolic reactions. In this paper, a full-length cDNA encoding CPR (designated as CmCPR) was characterized in the rice leaffolder, Cnaphalocrocis medinalis (Guenée), a serious lepidopteran rice pest. The complete open reading frame of CmCPR was 2046 bp, encoding a protein consists of 681 amino acid residues. The secondary structure of CmCPR protein showed marked features of classical CPRs such as N-terminal anchor, conserved functional domains, and catalytic residues. Phylogenic analysis showed that CmCPR was clustered together with CPRs from other lepidopteran species. Recombinant CmCPR protein was expressed in E. coli, and the activity and kinetic parameters of the enzyme were determined. Quantitative reverse transcription-PCR showed that the highest expression levels of CmCPR were detected in fourth- and fifth-instar larvae, and the transcriptional level in larval midgut tended to be higher than those in other tissues. Exposure to sublethal concentrations of three insecticides, abamectin, chlorpyrifos, and chlorantraniliprole, led to upregulated expression of CmCPR and several P450 genes. This work is the first report of molecular characterization of CPR gene in Cn. medinalis.






Similar content being viewed by others
References
Andersen JF, Utermohlen JG, Feyereisen R (1994) Expression of housefly CYP6A1 and NADPH–cytochrome P450 reductase in Escherichia coli and reconstitution of an insecticide-metabolizing P450 system. Biochemistry 33:2171–2177
Camacho C, Coulouris G, Avagyan V, Ma N, Papadopoulos J, Bealer K et al (2009) BLAST+: architecture and applications. BMC Bioinform 10:421
Chen X, Zhang Y (2015) Identification and characterization of NADPH-dependent cytochrome P450 reductase gene and cytochrome b 5 gene from Plutella xylostella: possible involvement in resistance to beta-cypermethrin. Gene 558:208–214
Feyereisen R (1999) Insect P450 enzymes. Ann Rev Entomol 44:507–533
Girhard M, Tieves F, Weber E, Smit M, Urlacher V (2013) Cytochrome P450 reductase from Candida apicola: versatile redox partner for bacterial P450s. Appl Microbiol Biotechnol 97:1625–1635
Hatano R, Scott JG (1993) Anti-P450lpr antiserum inhibits the activation of chlorpyrifos to chlorpyrifos oxon in house fly microsomes. Pestic Biochem Physiol 45:228–233
Horike N, Takemori H, Nonaka Y, Sonobe H, Okamoto M (2000) Molecular cloning of NADPH–cytochrome P450 oxidoreductase from silkworm eggs. Eur J Biochem 267:6914–6920
Hovemann BT, Sehlmeyer F, Malz J (1997) Drosophila melanogaster NADPH–cytochrome P450 oxidoreductase: pronounced expression in antennae may be related to odorant clearance. Gene 189:213–219
Hu Z, Lin Q, Chen H, Li Z, Yin F, Feng X (2014) Identification of a novel cytochrome P450 gene, CYP321E1 from the diamondback moth, Plutella xylostella (L.) and RNA interference to evaluate its role in chlorantraniliprole resistance. Bull Entomol Res 104:716–723
Huang Y, Lu X-P, Wang L-L, Wei D, Feng Z-J, Zhang Q et al (2015) Functional characterization of NADPH–cytochrome P450 reductase from Bactrocera dorsalis: possible involvement in susceptibility to malathion. Sci Rep 5:18394
Kaewpa D, Boonsuepsakul S, Rongnoparut P (2007) Functional expression of mosquito NADPH–cytochrome P450 reductase in Escherichia coli. J Econ Entomol 100:946–953
Kawazu K, Shintani Y, Tatsuki S (2014) Effect of multiple mating on the reproductive performance of the rice leaffolder moth, Cnaphalocrocis medinalis (Lepidoptera: Crambidae). Appl Entomol Zool 49:519–524
Lamb DC, Warrilow AGS, Venkateswarlu K, Kelly DE, Kelly SL (2001) Activities and kinetic mechanisms of native and soluble NADPH–cytochrome P450 reductase. Biochem Biophys Res Commun 286:48–54
Lamb DC, Kim Y, Yermalitskaya LV, Yermalitsky VN, Lepesheva GI, Kelly SL et al (2006) A second FMN binding site in yeast NADPH–cytochrome P450 reductase suggests a mechanism of electron transfer by diflavin reductases. Structure 14:51–61
Li X, Schuler MA, Berenbaum MR (2007) Molecular mechanisms of metabolic resistance to synthetic and natural xenobiotics. Ann Rev Entomol 52:231–253
Li S-W, Yang H, Liu Y-F, Liao Q-R, Du J, Jin D-C (2012) Transcriptome and gene expression analysis of the rice leaf folder, Cnaphalocrosis medinalis. PLoS One 7:e47401
Lian LY, Widdowson P, McLaughlin LA, Paine MJI (2011) Biochemical comparison of Anopheles gambiae and human NADPH P450 reductases reveals different 2′-5′-ADP and FMN binding traits. PLoS One 6:e20574
Liu S, Liang Q-M, Huang Y-J, Yuan X, Zhou W-W, Qiao F et al (2013) Cloning, functional characterization, and expression profiles of NADPH–cytochrome P450 reductase gene from the Asiatic rice striped stem borer, Chilo suppressalis (Lepidoptera: Pyralidae). Comp Biochem Physiol B 166:225–231
Liu D, Zhou X, Li M, Zhu S, Qiu X (2014) Characterization of NADPH–cytochrome P450 reductase gene from the cotton bollworm, Helicoverpa armigera. Gene 545:262–270
Liu S, Liang Q-M, Zhou W-W, Jiang Y-D, Zhu Q-Z, Yu H et al (2015a) RNA interference of NADPH–cytochrome P450 reductase of the rice brown planthopper, Nilaparvata lugens, increases susceptibility to insecticides. Pest Manag Sci 71:32–39
Liu S, Rao X-J, Li M-Y, Li S-G (2015b) Identification and expression profiles of putative cytochrome P450 monooxygenase genes from Cnaphalocrocis medinalis (Lepidoptera: Pyralidae). Entomol Res 45:141–149
Lycett GJ, McLaughlin LA, Ranson H, Hemingway J, Kafatos FC, Loukeris TG et al (2006) Anopheles gambiae P450 reductase is highly expressed in oenocytes and in vivo knockdown increases permethrin susceptibility. Insect Mol Biol 15:321–327
Maïbèche-Coisne M, Merlin C, François M-C, Porcheron P, Jacquin-Joly E (2005) P450 and P450 reductase cDNAs from the moth Mamestra brassicae: cloning and expression patterns in male antennae. Gene 346:195–203
Murataliev MB, Ariño A, Guzov VM, Feyereisen R (1999) Kinetic mechanism of cytochrome P450 reductase from the house fly (Musca domestica). Insect Biochem Mol Biol 29:233–242
Park S, Kim Y-S, Rupasinghe SG, Schuler MA, Back K (2013) Rice P450 reductases differentially affect P450-mediated metabolism in bacterial expression systems. Bioprocess Biosyst Eng 36:325–331
Pfaffl MW (2001) A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res 29:e45
Qiu Y, Tittiger C, Wicker-Thomas C, Le Goff G, Young S, Wajnberg E et al (2012) An insect-specific P450 oxidative decarbonylase for cuticular hydrocarbon biosynthesis. Proc Natl Acad Sci USA 109:14858–14863
Riga M, Tsakireli D, Ilias A, Morou E, Myridakis A, Stephanou EG et al (2014) Abamectin is metabolized by CYP392A16, a cytochrome P450 associated with high levels of acaricide resistance in Tetranychus urticae. Insect Biochem Mol Biol 46:43–53
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729
Tang Q-Y, Zhang C-X (2013) Data processing system (DPS) software with experimental design, statistical analysis and data mining developed for use in entomological research. Insect Sci 20:254–260
Wang M, Roberts DL, Paschke R, Shea TM, Masters BSS, Kim J-JP (1997) Three-dimensional structure of NADPH–cytochrome P450 reductase: prototype for FMN- and FAD-containing enzymes. Proc Natl Acad Sci USA 94:8411–8416
Wang K, Peng X, Zuo Y, Li Y, Chen M (2016) Molecular cloning, expression pattern and polymorphisms of NADPH-cytochrome P450 reductase in the bird cherry-oat aphid Rhopalosiphum padi (L.). PLoS One 11:e0154633
Wen Z, Pan L, Berenbaum MR, Schuler MA (2003) Metabolism of linear and angular furanocoumarins by Papilio polyxenes CYP6B1 co-expressed with NADPH cytochrome P450 reductase. Insect Biochem Mol Biol 33:937–947
Xia C, Hamdane D, Shen AL, Choi V, Kasper CB, Pearl NM et al (2011) Conformational changes of NADPH-cytochrome P450 oxidoreductase are essential for catalysis and cofactor binding. J Biol Chem 286:16246–16260
Yang C-Q, Lu S, Mao Y-B, Wang L-J, Chen X-Y (2010) Characterization of two NADPH: cytochrome P450 reductases from cotton (Gossypium hirsutum). Phytochemistry 71:27–35
Zhang S-K, Ren X-B, Wang Y-C, Su J (2014) Resistance in Cnaphalocrocis medinalis (Lepidoptera: Pyralidae) to new chemistry insecticides. J Econ Entomol 107:815–820
Zhang Y, Wang Y, Wang L, Yao J, Guo H, Fang J (2016) Knockdown of NADPH-cytochrome P450 reductase results in reduced resistance to buprofezin in the small brown planthopper, Laodelphax striatellus (fallén). Pestic Biochem Physiol 127:21–27
Zhao C, Tang T, Feng X, Qiu L (2014) Cloning and characterisation of NADPH-dependent cytochrome P450 reductase gene in the cotton bollworm, Helicoverpa armigera. Pest Manag Sci 70:130–139
Zhao C, Feng X, Tang T, Qiu L (2015) Isolation and expression analysis of CYP9A11 and cytochrome P450 reductase gene in the beet armyworm (Lepidoptera: Noctuidae). J Insect Sci 15:122
Zheng X, Ren X, Su J (2011) Insecticide susceptibility of Cnaphalocrocis medinalis (Lepidoptera: Pyralidae) in China. J Econ Entomol 104:653–658
Zhou X, Li M, Sheng C, Qiu X (2011) NADPH-cytochrome P450 oxidoreductase from the chicken (Gallus gallus): sequence characterization, functional expression and kinetic study. Comp Biochem Physiol C 153:53–59
Zhu F, Sams S, Moural T, Haynes KF, Potter MF, Palli SR (2012) RNA interference of NADPH-cytochrome P450 reductase results in reduced insecticide resistance in the bed bug, Cimex lectularius. PLoS One 7:e31037
Acknowledgements
This work was supported by the National Natural Science Foundation of China (31401734), the National Key Research and Development Program of China (2016YFD0200205-7), and the Anhui Provincial Natural Science Foundation (1708085QC50).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhang, YX., Li, SG., Rao, XJ. et al. Molecular characterization of a NADPH–cytochrome P450 reductase gene from the rice leaffolder, Cnaphalocrocis medinalis (Lepidoptera: Pyralidae). Appl Entomol Zool 53, 19–27 (2018). https://doi.org/10.1007/s13355-017-0523-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13355-017-0523-y