Abstract
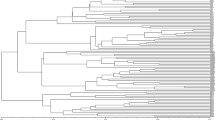
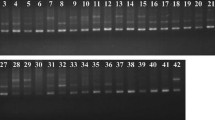
Mulberry, Morus spp. has a wide range of use, the chief among them is to feed the silk producing caterpillar Bombyx mori L. (Bombycidae; Bombycoidea). As a homeland of mulberry, India has a number of indigenous mulberry species, of which a few are widely cultivated. In the present investigation genetic distance among such eleven mulberry cultivars originated from six different states of India covering a wide geographic area extending from 15° N to 32° N latitude and 72°E to 89°E longitude was studied using inter-simple sequence repeat primers. Out of the 20primers tested, 13 primers, viz, nine di-nucleotide, three tri-nucleotide and one penta-nucleotide repeats, gave clear and reproducible band profiles. While the (AT)n rich primers could not amplify the DNA, the (GA)n, (AC)n and (AG)n rich primers gave excellent amplification profiles. The genetic distance among the cultivars varied from a minimum of 0.053, between Punjab local and Bombay local, to a maximum of 0.431, between Almora local andSujanpur-5. Clustering of the cultivars according to nearest neighbor method created three groups. The north-Indian cultivars made a separate and distinct group while the cultivars originated from eastern and southern India occupied a distinct position. Almora local was found quite different from others. The first two canonical functions identified through discriminant function analysis accounted for 91.2% of the total variability. Distribution of cultivars belonging to six different zones on canonical matrix realized from Discriminant Function Analysis (DFA) revealed wider variability for West Bengal, Karnataka and Punjab which reaches the group centroids of Uttar Pradesh and Himachal Pradesh. This attests to the past contribution of West Bengal in east and Karnataka in south towards development of mulberry cultivars indifferent parts of India. Step-wise linear regression analysis, further, identified two markers (825.1400 and835.750) associated with leaf yield, which also satisfied the Beta estimation, thereby testifying strong association of these two markers with leaf yield. This finding along with the classification of the eleven cultivars bear strong relevance to mulberry breeding for different agro climatic areas.
Similar content being viewed by others
References
Agaki, H., Y. Yokozeki, A. Inagaki, A. Nakamura & T. Fugimura, 1996. A codominant DNA marker closely linked to the rice nuclear restorer gene, Rf-1, identified with inter-SSR fingerprinting. Genome 39: 1205-1209.
Breto, M.P., C. Ruiz, J.A. Pina & M.J. Asins, 2001. The diversification of Citrus clementina Hort. Ex Tan., a vegetatively propagated crop species. Mol Phylogenet Evol 21: 285-293.
Chatterjee, S.N., 1993. Silkworm breeding in India. Sericologia 33: 427-447.
Culley, T.M. & A.D. Wolfe, 2001. Population genetic structure of the cleistogamous plant species Viola pubescens Aiton (Violaceae), as indicated by allozyme and ISSR molecular markers. Heridity 86 (Pt 5): 545-556.
Dandin, S.B., R. Kumar, S. Rabindran & M.S. Jolly, 1987. Crossability studies in mulberry. Indian J Sericult 26: 11-14.
Das, B.C. & S. Krishnaswami, 1965. Some observations on interspecific hybridization in mulberry. Indian J Sericulte 4: 1-8.
Deshpande, A.U., G.S. Apte, R.A. Bahulikar, M.D. Lagu, B.G. Kulkarni, H.S. Suresh, N.P. Singh, M.K. Rao, V.S. Gupta, A. Pant & P.K. Rnjekar, 2001. Genetic diversity across natural populations of montane plant species from the Western Ghats, India reveled by intersimple sequence repeats. Mol Ecol 10: 2397-2408.
Devi, S.P. & A. Agarwal, 1989. Scanning of early sprouting varieties of mulberry. Sci & Cult (India) 54: 87-88.
Devos, K.M. & M.D. Gale, 1992. The use of random amplified polymorphic DNA markers in wheat. Theor Appl Genet 89: 567-572.
Fang, D.Q. & M.L. Roose, 1997. Identification of closely related citrus cultivars with inter-simple sequence repeat markers. Theor Appl Genet 95: 408-417.
Fang, D.Q., M.L. Roose, R.R. Krueger & Federici, 1997. Finger printing trifoliate orange germplasm accessions with isozymes, RFLP and inter-simple sequence repeat markers. Theor Appl Genet 95: 211-219.
Felsenstein, J., 1993. PHYLIP (Phylogeny inference package) version 3.5c. Distributed by the author. Department of Genetics, University of Washington, Seattle.
Fotedar, R.K. & S.B. Dandin, 1998. Genetic divergence in the mulberry. Sericologia 38: 115-125.
Frie, O.M., C.W. Stuber & M.M. Goodman, 1986. Use of allozymes as genetic markers for predicting genetic performance in maize single cross hybrids. Crop Science 26: 37-42.
Grafius, J.E., 1956. Components of yield in oat: a geaometrical interpretation. Agron J 48: 419-423.
He, G.H., L. Hou, D.M. Li, X.Y. Luo, G.Q. Niu, M. Tang & Y. Pei, 2002. Prediction of yield and yield components in hybrid rice by using molecular markers. Yi Chuan Xue Bao 29: 438-444.
Hirano, H. & K. Naganuma, 1979. Inheritance of peroxidase isozyme in mulberry (morus spp.). Euphytica 28: 73-79.
Hotta, T., 1954 Taxonomical Study on the Cultivated Mulberry in Japan. 125 faculty of textile fibres Kyoto Univ. Industrial art and textile fibres, Kyoto.
Kantety, R.V., X. Zhang, J.L. Bennetzen J.L. & B.Z. Zehr, 1995. Assessment of genetic diversity in dent and popcorn (Zea mays L.) inbred lines using ISST-simple sequence repeat (ISSR) amplification. Mol Breed 1: 365-373.
Katsumata, T., 1972. Mulberry species in West Jawa and their peculiarities. J Seric Sci Japan 42: 213-223.
Kitaurak, 1983. Morphological and biochemical differences among four mulberry species from Java. J Seric Sci Japan 52: 198-202.
Koidzumi, G., 1917. Taxonomy and phytogeography of the genus Morus. Bull Seric Exp Station, Tokyo (Japan) 3: 1-62.
Linneaus, C., 1753. Species plantarum: 986.
Lou, C.F., Y.Z. Zhang & J.M. Zhou, 1998. Polymorphisms of genomic DNA in parents and their resulting hybrids in mulberry (Morus). Sericologia 38: 437-445.
Mala, V.R., H.K. Chaturvedi & A. Sarkar, 1997. Multivaraite analysis as an aid to genotype selection for breeding in mulberry. Indian J Seric 36: 111-115.
Nagaoka, T. & Y. Ogihara, 1997. Applicability of inter-simple sequence repeat polymorphism in wheat for use as DNA markers in comparison to RFLP and RAPD markers. Theor Appl Genet 94: 597-602.
Nei, M. & H. Li W, 1979. Mathematical model for study the genetic variation in terms of restriction endonucleases. Proc Natl Acad Sci USA 74: 5267-5273.
Press, S.J. & S. Wilson, 1978. Choosing between logistic regression and discriminant analysis. J Amer Stat Assoc 73: 699-705.
Provost A. & M.J. Wilkinson, 1999. A newsystem of comparinmg PCR primers applied to ISSR fingerprinting of potato cultivars. Theor Appl Genet 98: 107-112.
Rafalski, J.A., S.V. Tingey & J.G.K. Williams, 1991. RAPD markers-a new technology for genetic mapping and plant breeding. AgBiotech News & Info 3: 645-648.
Raina, S.N., V. Rani, T. Kojima, Y. Ogihara, K.P. Singh & R.M. Devarumath, 2001. RAPD and ISSR finger prints as useful genetic markers for analysis of genetic diveristy, Varietal identification, and phylogenetic relationships in parent (Arachis hypogea) cultivars and wild species. Genome 44: 763-772.
Ratnaparkhe, M.B., D.K. Santra, A. Tullu & F.J. Muehlbauer, 1998. Inheritance of inter-simple repeat polymorphism and linkage with a fusarium wilt resistance gene in chickpea. Theor Appl Genet 96: 348-353.
Ravindran, S.V., V. Anand Rao, V. Girish Naik, A. Tikader, P. Mukherjee & K. Thengavelu, 1997. Distribution and variation in mulberry germplasm. Indian J Pl Genet Res 10(2): 233-242.
Sau, H., P.K. Sahu, B.R.D. Yadav & B. Saratchandra, 1995 Evaluation of mulberry (Morus spp) genetic resources-I. Sprouting, survival and rooting ability. J Envir Res (India) 3: 11-13.
Sethuraman, B.N., T.P. Mohandas & S.N. Chatterjee, 2002. DNA fingerprinting with homologous multilocus probes and search for DNA markers associated with yield attributes in silkworm, Bombyx mori. Eur J Entomol 99: 267-276.
Simith, J.S.C., 1984. Genetic variabilty within US hybrid maize: multivariate analysis of isozyme data. Cro Science 24: 1041-1046.
Tikader, A., M. Shamsuddin, K. Vijayan & T. Pavankumar 1995. Survival potential in different varieties of mulberry (Morus species) Indian J Agric Sci 65: 133-135.
Tsumura, Y., K. Ohba & S.H. Strauss, 1996. Diversity and inheritance of Inter-simple repeat polymorphism in Douglas for (Pseudotsuga menziesii) and Sugi (Crypromeria japonica). Theor Appl Genet 92: 40-45.
Vavilov, N.I., 1951 The origin, immunity and breeding of cultivated plants. Translated from Russian by K.S. Chester. Chrenica Botanica Vv 1: 6.
Venkateswarlu, M., B.N. Susheelamma, N. Suryanarayana, N.K. Dwivedi & K. Sengupta, 1989. Peroxidase isozyme banding patterns in Aneuploids of mulberry. Sericologia 29: 99-104.
Vijayan, K., A. Tikader, S.P. Chkraborti, B.N. Roy & T. Pavankumar, 1994. Studies on Crossability of Mulberry in Polyploid Breeding. 2nd National seminar on prospects of Sericulture in India, University of Madras, Vellore, TamilNadu, India, March 7-9, pp. 79-85.
Vijayan, K., K.K. Das, S.G. Doss, S.P. Chakraborti & B.N. Roy, 1999. Genetic divergence in indigenous mulberry (Morus spp.) genotypes. Indian J Agric Sci 69: 851-853.
Wang, Z., J.L. Weber, G. Zhong & S.D. Tanksley, 1994. Survey of plant short tandem DNA repeats. Theor Appl Genet 88: 1-6.
Weiguo, Z., P. Yile & H. Minren, 2000. RAPD analysis for the germplasm resources of the genus mulberry, Morus L. Canye Kexue 26(4): 197-204.
Wu, Y.Y., T.Z. Zhang & J.M. Yin, 2001. Genetic diversity detected by DNA markers and phenotypes in Upland cotton. YI Chuan Xue Bao (China) 28: 1040-1050.
Yonash, N., E.D. Heller, J. Hillel & A. Cahaner, 2000. Detection of RFLP markers associated with antibody response in meattype chickens Haplotype/genotype, single-band and multiband analysis of RFLP in the major histocompatibility complex. J Heredity 91: 24-30.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Vijayan, K., Chatterjee, S.N. ISSR profiling of Indian cultivars of mulberry (Morus spp.) and its relevance to breeding programs. Euphytica 131, 53–63 (2003). https://doi.org/10.1023/A:1023098908110
Issue Date:
DOI: https://doi.org/10.1023/A:1023098908110