Abstract
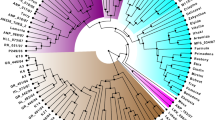
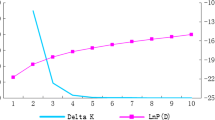
Estimation of genetic diversity and relative relatedness in breeding materials is critical for improving breeding efficiency. To compare the ability of single nucleotide polymorphism (SNP) and insertion-deletion (InDel) markers for characterizing cultivated tomato germplasm, 120 SNPs and 109 InDels were used to genotype 191 tomato inbred lines representing cherry tomato, traditional varieties, and contemporary lines. The results showed that SNPs provided more information on genetic diversity than the InDels. The expected heterozygosity (He) of SNPs and InDels averaged 0.384 and 0.265, respectively, and the polymorphic information content (PIC) of these two markers was 0.302 and 0.221, respectively. Except for the cherry tomato group, the traditional group showed higher He and PIC for the SNP data, and the contemporary group had the higher InDel diversity. Population structure analysis revealed that the traditional varieties constituted distinct subpopulations relative to the contemporary lines with both marker systems, and three subpopulations were found within the traditional group with SNPs. Additionally, SNPs provided more resolution in discriminating the closely related tomato lines, and InDels may be more effective at resolving genotypes from an inter-gene pool. A lower correlation (R = 0.4155) was found between SNPs and InDels based on the genetic distances among accessions. The present study systematically compares the performance of SNP and InDel markers for population genetics analysis in cultivated tomato. These results contribute to the choice of molecular marker type for analysis of genetic diversity and other genetic studies in tomato.



Similar content being viewed by others
REFERENCES
Menda, N., Semel, Y., Peled, D., et al., In silico screening of a saturated mutation library of tomato, Plant J., 2004, vol. 38, no. 5, pp. 861—872.
Shirasawa, K. and Hirakawa, H., DNA marker applications to molecular genetics and genomics in tomato, Breed. Sci., 2013, vol. 63, no. 1, pp. 21—30.
Archak, S., Karihaloo J.L., and Jain, A., RAPD markers reveal narrowing genetic base of Indian tomato cultivars, Curr. Sci., 2002, vol. 82, no. 9, pp. 1139—1143.
Mazzucato, A., Papa, R., Bitocchi, E., et al., Genetic diversity, structure and marker—trait associations in a collection of Italian tomato (Solanum lycopersicum L.) landraces, Theor. Appl. Genet., 2008, vol. 116, no. 5, pp. 657—669.
Sim, S.C., Van Deynze, A., Stoffel, K., et al., High-density SNP genotyping of tomato (Solanum lycopersicum L.) reveals patterns of genetic variation due to breeding, PLoS One, 2012, vol. 7, no. 9. e45520.
Corrado, G., Piffanelli, P., Caramante, M., et al., SNP genotyping reveals genetic diversity between cultivated landraces and contemporary varieties of tomato, BMC Genomics, 2013, vol. 14, p. 835.
Miller, J.C. and Tanksley, S.D., RFLP analysis of phylogenetic relationships and genetic variation in the genus, Theor. Appl. Genet., 1990, vol. 80, no. 4, pp. 437—448.
Park, Y.H., West, M.A., and St Clair, D.A., Evaluation of AFLPs for germplasm fingerprinting and assessment of genetic diversity in cultivars of tomato (Lycopersicon esculentum L.), Genome, 2004, vol. 47, no. 3, pp. 510—518.
Ranc, N., Munos, S., Santoni, S., et al., A clarified position for Solanum lycopersicum var. cerasiforme in the evolutionary history of tomatoes (Solanaceae), BMC Plant Biol., 2008, vol. 8, p. 130.
Chen, J., Wang, H., Shen, H., et al., Genetic variation in tomato populations from four breeding programs revealed by single nucleotide polymorphism and simple sequence repeat markers, Sci. Hortic., 2009, vol. 122, no. 1, pp. 6—16.
Bai, Y. and Lindhout, P., Domestication and breeding of tomatoes: what have we gained and what can we gain in the future?, Ann. Bot., 2007, vol. 100, no. 5, pp. 1085—1094.
Frary, A., Xu, Y., Liu, J., et al., Development of a set of PCR-based anchor markers encompassing the tomato genome and evaluation of their usefulness for genetics and breeding experiments, Theor. Appl. Genet., 2005, vol. 111, no. 2, pp. 291—312.
Tomato Genome Consortium, The tomato genome sequence provides insights into fleshy fruit evolution, Nature, 2012, vol. 485, no. 7400, pp. 635—641.
Yang, W., Bai, X., Kabelka, E., et al., Discovery of single nucleotide polymorphisms in Lycopersicon esculentum by computer aided analysis of expressed sequence tags, Mol. Breed., 2004, vol. 14, no. 1, pp. 21—34.
Jiménez Gómez, J.M. and Maloof, J.N., Sequence diversity in three tomato species: SNPs, markers, and molecular evolution, BMC Plant Biol., 2009, vol. 9, p. 85.
Sim, S.C., Robbins, M.D., Chilcott, C., et al., Oligonucleotide array discovery of polymorphisms in cultivated tomato (Solanum lycopersicum L.) reveals patterns of SNP variation associated with breeding, BMC Genomics, 2009, vol. 10, p. 466.
Hamilton, J.P., Sim, S.C., Stoffel, K., et al., Single nucleotide polymorphism discovery in cultivated tomato via sequencing by synthesis, Plant Genome, 2012, vol. 5, no. 1, pp. 17—29.
Lin, T., Zhu, G.T., Zhang, J.H., et al., Genomic analyses provide insights into the history of tomato breeding, Nat. Genet., 2014, vol. 46, no. 11, pp. 1220–1228.
Yang, J., Wang, Y., Shen, H., et al., In silico identification and experimental validation of insertion—deletion polymorphisms in tomato genome, DNA Res., 2014, vol. 21, no. 4, pp. 429—438.
Tam, S.M., Mhiri, C., Vogelaar, A., et al., Comparative analyses of genetic diversities within tomato and pepper collections detected by retrotransposon-based SSAP, AFLP and SSR, Theor. Appl. Genet., 2005, vol. 110, no. 5, pp. 819—831.
Frascaroli, E., Schrag, T.A., and Melchinger, A.E., Genetic diversity analysis of elite European maize (Zea mays L.) inbred lines using AFLP, SSR, and SNP markers reveals ascertainment bias for a subset of SNPs, Theor. Appl. Genet., 2013, vol. 126, no. 1, pp. 133—141.
Filippi, C.V., Aguirre, N., Rivas, J.G., et al., Population structure and genetic diversity characterization of a sunflower association mapping population using SSR and SNP markers, BMC Plant Biol., 2015, vol. 15, p. 52.
Müller, B.S.F., Pappas G.J. Jr., Valdisser, P.A.M.R., et al., An operational SNP panel integrated to SSR marker for the assessment of genetic diversity and population structure of the common bean, Plant Mol. Biol. Rep., 2015, vol. 33, no. 6, pp. 1697—1711.
Wang, T., Zou, Q.D., Qi, S.Y., et al., Analysis of genetic diversity and population structure in a tomato (Solanum lycopersicum L.) germplasm collection based on single nucleotide polymorphism markers, Genet. Mol. Res., 2016, vol. 15.
Shirasawa, K., Isobe, S., Hirakawa, H., et al., SNP discovery and linkage map construction in cultivated tomato, DNA Res., 2010, vol. 17, no. 6, pp. 381—391.
Van Deynze, A., Stoffel, K., Buell, C.R., et al., Diversity in conserved genes in tomato, BMC Genomics, 2007, vol. 8, p. 465.
Stewart, C.N., Jr. and Via, L.E., A rapid CTAB DNA isolation technique useful for RAPD fingerprinting and other PCR applications, Biotechniques, 1993, vol. 14, no. 5, pp. 748—750.
Chang, H.W., Cheng, Y.H., Chuang, L.Y., et al., SNP-RFLPing 2: an updated and integrated PCR-RFLP tool for SNP genotyping, BMC Bioinf., 2010, vol. 11, p. 173.
Ge, Y., Ramchiary, N., Wang, T., et al., Development and linkage mapping of unigene-derived microsatellite markers in Brassica rapa L., Breed. Sci., 2011, vol. 61, no. 2, pp. 160—167.
Peakall, R. and Smouse, P.E., GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research—an update, Bioinformatics, 2012, vol. 28, no. 19, pp. 2537—2539.
Liu, K. and Muse, S.V., PowerMarker: an integrated analysis environment for genetic marker analysis, Bioinformatics, 2005, vol. 21, no. 9, pp. 2128—2129.
Pritchard, J.K., Stephens, M. and Donnelly, P., Inference of population structure using multilocus genotype data, Genetics, 2000, vol. 155, pp. 945—959.
Evanno, G., Regnaut, S., and Goudet, J., Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study, Mol. Ecol., 2005, vol. 14, no. 8, pp. 2611—2620.
Corrado, G., Caramante, M., Piffanelli, P., et al., Genetic diversity in Italian tomato landraces: implications for the development of a core collection, Sci. Hortic., 2014, vol. 168, pp. 138—144.
Sim, S.C., Robbins, M.D., Van Deynze, A., et al., Population structure and genetic differentiation associated with breeding history and selection in tomato (Solanum lycopersicum L.), Heredity, 2011, vol. 106, no. 6, pp. 927—935.
Hamblin, M.T., Warburton, M.L., and Buckler, E.S., Empirical comparison of simple sequence repeats and single nucleotide polymorphisms in assessment of maize diversity and relatedness, PLoS One, 2007, vol. 12, no. 12. e1367.
Wurschum, T., Langer, S.M., Longin, C.F., et al., Population structure, genetic diversity and linkage disequilibrium in elite winter wheat assessed with SNP and SSR markers, Theor. Appl. Genet., 2013, vol. 126, no. 6, pp. 1477—1486.
Yang, X., Xu, Y., Shah, T., et al., Comparison of SSRs and SNPs in assessment of genetic relatedness in maize, Genetica, 2011, vol. 139, no. 8, pp. 1045—1054.
Rick, C.M. and Fobes, J.F., Allozyme variation in the cultivated tomato and closely related species, Bull. Torrey Bot. Club, 1975, vol. 102, no. 6, pp. 376—384.
Moragues, M., Comadran, J., Waugh, R., et al., Effects of ascertainment bias and marker number on estimations of barley diversity from high-throughput SNP genotype data, Theor. Appl. Genet., 2010, vol. 120, no. 8, pp. 1525—1534.
Causse, M., Desplat, N., Pascual, L., et al., Whole genome resequencing in tomato reveals variation associated with introgression and breeding events, BMC Genomics, 2013, vol. 14, p. 791.
Varshney, R.K., Baum, M., Guo, P., et al., Features of SNP and SSR diversity in a set of ICARDA barley germplasm collection, Mol. Breed., 2010, vol. 26, no. 2, pp. 229—242.
Jones, E.S., Sullivan, H., Bhattramakki, D., et al., A comparison of simple sequence repeat and single nucleotide polymorphism marker technologies for the genotypic analysis of maize (Zea mays L.), Theor. Appl. Genet., 2007, vol. 115, no. 3, pp. 361—371.
Varshney, R.K., Thiel, T., Sretenovic Rajicic, T., et al., Identification and validation of a core set of informative genic SSR and SNP markers for assaying functional diversity in barley, Mol. Breed., 2007, vol. 22, no. 1, pp. 1—13.
Woodhead, M., Russell, J., Squirrell, J., et al., Comparative analysis of population genetic structure in Athyrium distentifolium (Pteridophyta) using AFLPs and SSRs from anonymous and transcribed gene regions, Mol. Ecol., 2010, vol. 14, no. 6, pp. 1681—1695.
ACKNOWLEDGMENTS
This work was supported by the National Key R&D Program of China (2017YFD0101902).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
The authors declare that they have no conflict of interest. This article does not contain any studies involving animals or human participants performed by any of the authors.
Additional information
The article is published in the original.
Supplementary material
Rights and permissions
About this article
Cite this article
Wang, T., Li, H.T., Zhu, H. et al. Comparative Analyses of Genetic Variation in a Tomato (Solanum lycopersicum L.) Germplasm Collection with Single Nucleotide Polymorphism and Insertion-Deletion Markers. Russ J Genet 55, 204–211 (2019). https://doi.org/10.1134/S1022795419020182
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S1022795419020182