Abstract
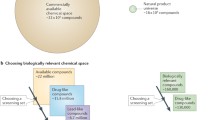
Genomic-based methodologies are increasingly used at all stages of drug development. The most extensive applications have occurred in early drug discovery stages due to advances in technologies that allow for automated synthesis and characterization of organic compounds, and for high-throughput screening of these molecules against known drug targets. The adaptation of genomic-based methodologies in later stages of drug development presents a more difficult task.
In this review we describe how genomics can be used to identify previously uncharacterized pharmacologic actions that provide a basis for the development of new classes of antimycotic agents or for adverse event aversion. Clinically, novel antimycotics are gravely needed. This review provides a perspective on new technologies that will bridge the gap between drug discovery and development that may enable more rapid access to new antimycotic agents.





Similar content being viewed by others
References
Shlaes DM, Progan SJ, Edwards JE. Antibiotic discovery: state of the state. Am Soc Microbiol News 2004; 70: 275–81
Andriole VT. Current and future antifungal therapy: new targets for antifungal agents. J Antimicrob Chemother 1999; 44: 151–62
Pfaller M, Wenzel R. Impact of the changing epidemiology of fungal infections in the 1990s. Eur J Clin Microbiol Infect Dis 1992; 11: 287
Lupetti A, Danesi R, Campa M, et al. Molecular basis of resistance to azole antifungals. Trends Mol Med 2002; 8(2): 76–81
Pacetti SA, Gelone SP. Caspofungin acetate for treatment of invasive fungal infections. Ann Pharmacother 2003; 37(1): 90–8
Schultes R, Raffauf R. The healing forest: medicinal and toxic plants of the Northwest Amazonia. Portland (OR): Discorides Press, 1990
Faulkner DJ. Marine natural products. Nat Prod Reps 2001; 18: 1–49
Urban S, Hickford SJH, Blunt JW, et al. Bioactive marine alkaloids. Current Org Chem 2000; 4: 765–807
Newman DJ, Cragg GM, Snader KM. Natural products as sources of new drugs over the period 1981–2002. J Nat Prod 2003; 66: 1022–37
Cragg GM, Newman DJ, Snader KM. Natural products in drug discovery and development. J Nat Prod 1997; 60: 52–60
Vicente MF, Basilio A, Cabello A, et al. Microbial natural products as a source of antifungals. Clin Microbiol Infect 2003; 9: 15
Kinghorn AD, Balandrin MF, editors. Human medicinal agents from plants. ACS Symposium Series 534. Washington, DC: American Chemical Society, 1993: 2–12
Cowan MM. Plant products as antimicrobial agents. Clin Microbiol Rev 1999; 12: 564–82
Graybill JR. The future of antifungal therapy. Clin Infect Dis 1996; 22Suppl. 2: S166–78
Denning DW. Echinocandins and pneumocandins: a new antifungal class with a novel mode of action. J Antimicrob Chemother 1997; 40: 611–4
Sawistowska-Schroder ET, Kerridge D, Perry H. Echinocandin inhibition of 1,3-beta-D-glucan synthase from Candida albicans. FEBS Lett 1984; 173: 134
Singh SB, Zink DL, Doss GA, et al. Citrafungins A and B, two new fungal metabolite inhibitors of GGTase I with antifungal activity. Org Lett 2004; 6: 337–40
Laakso JA, Raulli R, McElhaney-Feser GE, et al. CT2108A and B: new fatty acid synthase inhibitors as antifungal agents. J Nat Prod 2003; 66(8): 1041–6
Shibazaki M, Taniguchi M, Yokoi T, et al. YM-215343, a novel antifungal compound from Phoma sp. QN04621. J Antibiotics (Tokyo) 2004; (6): 379–82
Ma G, Khan SI, Jacob MR, et al. Antimicrobial and antileishmanial activities of hypocrellins A and B. Antimicrob Agents Chemother 2004; 48(11): 4450–2
El Sohly HN, Joshi AS, Nimrod AC, et al. Antifungal chalcones from Maclura tinctoria. Planta Med 2001; 67: 87–9
Li XC, Joshi AS, El Sohly HN, et al. Fatty acid synthase inhibitors from plants: isolation, structure elucidation, and SAR studies. J Nat Prod 2002; 65: 1909–14
Muhammad I, Dunbar DC, Takamatsu S, et al. Antimalarial, cytotoxic and antifungal alkaloids from Duguetia hadrantha. J Nat Prod 2001; 64: 559–62
El Sayed KA, Kelly M, Kara UAK, et al. New manzamine alkaloids with potent activity against infectious disease. J Am Chem Soc 2001; 123: 1804–8
Dunbar DC, Rimoldi JM, Clark AM, et al. Anti-cryptococcal and nitric oxide synthase inhibitory imidazole alkaloids from the calcareous sponge Leucetta cf chagosensis. Tetrahedron 2000; 56: 8795–8
Li XC, Dunbar DC, El Sohly HN, et al. A new naphthopyrone derivative from Cassia quinquangulata and structure revision of quinquangulin and its glycosides. J Nat Prod 2001; 64: 1153–6
Bedir E, Khan IA, Walker LA. Biologically active steroidal glycosides from Tribulus terrestris. Pharmazie 2002; 57: 491–4933
Zhang Z, ElSohly HN, Jacob MR, et al. New sesquiterpenoids from the root of Guatteria multivenia. J Nat Prod 2002; 65: 856–9
Liu S-C, Oguntimein BO, Hufford CD, et al. 3-methoxysampangine, a novel antifungal copyrine akaloid from Cleistopholis patens. Antimicrob Agents Chemother 1990; 34: 529–34
Clark AM. Sampangine and derivatives useful as an antifungal agent. Unversity of MIssissippi. US Patent 5,128,344.1992 July 7
Peterson JR, Zjawiony JK, Liu S-C, et al. Copyrine alkaloids: synthesis, spectroscopic characterization and antifungal/antimycobacterial activity of A- and B-ring functionalized sampangines. J Med Chem 1992; 35: 4069–77
Orabi KY, Li E, Clark AM, et al. Microbial transformation of sampangine. J Nat Prod 1999; 62: 988–92
Khan SI, Nimrod AC, Nitiss JL, et al. Antifungal activity of eupolauridine and its action on DNA topoisomerases. Antimicrob Agents Chemother 2002; 46: 1785–92
Hufford CD, Liu S-C, Clark AM, et al. Anticandidal activity of eupolauridine and onychine, alkaloids from Clesitopholis patens. J Nat Prod 1987; 50: 961
Clark AM. Antimicrobial compound [Eupolauridine] and compositions particularly effective against Candida albicans. US Patient 07/218,986. 1990 Oct
Li E, Clark AM, Hufford CD. Fungal evaluation of pseudolaric acid B, a major constituent of Pseudolarix kaempferi. J Nat Prod 1995; 58: 57–67
Onishi J, Meinz M, Thompson J, et al. Discovery of novel antifungal (1,3)-beta-D-glucan synthase inhibitors. Antimicrob Agents Chemother 2000; 44(2): 368–77
Cruz MC, Goldstein AL, Blankenship JR, et al. Calcineurin is essential for survival during membrane stress in Candida albicans. EMBO J 2002; 21(4): 546–59
Cruz MC, Goldstein AL, Blankenship J, et al. Rapamycin and less immunosuppressive analogs are toxic to Candida albicans and Cryptococcus neoformans via FKBP12-dependent inhibition of TOR. Antimicrob Agents Chemother 2001; 45(11): 3162–70
The SNP Consortium Ltd. Single nucleotide polymorphisms for biomedical research [online]. Available from URL: http://snp.cshl.org [Accessed 2005 Oct 10]
The Wellcome Trust. The human genome [online]. Available from URL: http://www.wellcome.ac.uk/en/genome/index.html [Accessed 2005 Oct 10]
Eisen MB, Spellman PT, Brown PO, et al. Cluster analysis and display of genome-wide expression patterns. Proc Nat Acad Sci 1998; 95: 14863–8
Tamayo P, Slonim D, Mesirov J, et al. Interpreting patterns of gene expression with self-organizing maps: methods and applications to hematopoietic differentiation. Proc Nat Acad Sci 1999; 96: 2907–12
Slonim DK, Tamayo P, Nesirov JP, et al. Class prediction and discovery using gene expression data. Information Systems 2003; 28(4): 243–268 [online]. Available from URL: http://www.broad.mit.edu/mpr/publications/projects/Leukemia/Slonim_et_al_2000.pdf [Accessed 2005 Oct 6]
Brown MPS, Grundy WN, Lin D, et al. Knowledge-based analysis of microarray gene expression data by using support vector machines. Proc Nat Acad Sci 2000; 97: 262–7
Sokal RR, Michener CD. A statistical method for evaluating systematic relationships. Univ Kans Sci Bull 1958; 38: 1409–38
Prigneau O, Porta A, Maresca B. Candida albicans CTN gene family is induced during macrophage infection: homology, disruption and phenotypic analysis of CTN3 gene. Fungal Genet Biol 2004; 41(8): 783–93
Beckerman J, Chibana H, Turner J, et al. Single-copy IMH3 allele is sufficient to confer resistance to mycophenolic acid in Candida albicans and to mediate transformation of clinical Candida species. Infect Immun 2001; 69(1): 108–14
Nakayama H, Mio T, Nagahashi S, et al. Tetracycline-regulatable system to tightly control gene expression in the pathogenic fungus Candida albicans. Infect Immun 2000; 68(12): 6712–9
Agarwal AK, Rogers PD, Baerson SR, et al. Genome-wide expression profiling of the response to polyene, pryimidine, azole and echinocandin anti-fungal agents in Saccharomyces cerevisiae. J Biol Chem 2003; 278: 34998–5015
Garrod AE. The incidence of alkaptonuria: a study in chemical individuality. 1902. Mol Med 1996; 2(3): 274–82
US Department of Health and Human Services. Guidance for industry: pharmacogenomic data submissions. March 2005 [online]. Available from URL: http://www.fda.gov/cder/guidance/6400fnl.pdf [Accessed 2005 Oct 10]
Butler D. Epidemiology set to get fast-track treatment. Nature 2001; 414: 38–43
Clifford RJ, Edmonson MN, Nguyen C, et al. Bioinformatics tools for single nucleotide polymorphism discovery and analysis. Ann N Y Acad Sci 2004; 1020: 101–9
Vogel F. Moderne probleme de humangenetik. Ergeb Inn Med Kinderheilkd 1959; 12: 52–125
Nebert DW. Extreme discordant phenotype methodology: an intuitive approach to clinical pharmacogenetics. Eur J Pharmacol 2000; 410: 107–20
Vesell ES. Pharmacogenetic perspectives gained from twin and family studies. Pharmacol Ther 1989; 41(3): 535–52
Tucker GT, Houston JB, Huang S-M. Optimizing drug development: strategies to assess drug metabolism/transporter interaction potential: toward a consensus. Clin Pharmacol Ther 2001; 70(2): 103–14
Bjornsson TD, Callaghan JT, Einolf HJ, et al. Pharmaceutical Research and Manufacturers of America (PhRMA) Drug Metabolism/Clinical Pharmacology Technical Working Group; FDA Center for Drug Evaluation and Research (CDER). The conduct of in vitro and in vivo drug-drug interaction studies: a Pharmaceutical Research and Manufacturers of America (PhRMA) perspective. Drug Metab Dispos 2003; 31(7): 815–32
Donato MT, Castell JV. Strategies and molecular probes to investigate the role of cytochrome P450 in drug metabolism. Clin Pharmacokinet 2003; 42(2): 153–78
Daly AK. Pharmacogenetics of the major polymorphic metabolizing enzymes. Fund Clin Pharmacol 2003; 17: 27–41
Sachse C, Brockmoller J, Bauer S, et al. Cytochrome P450 2D6 variants in a Caucasian population: allele frequencies and phenotypic consequences. Am J Hum Genet 1997; 60: 284–95
Vickers AEM, Sinclair JR, Zollinger M, et al. Multiple cytochrome P-450s involved in the metabolism of terbinafine suggest a limited potential for drug-drug interactions. Drug Metab Disp 1999; 27(9): 1029–38
Abdel-Rahman SM, Marcucci K, Boge T, et al. Potent inhibition of cytochrome P-450 2D6-mediated dextromethorphan O-demethylation by terbinafine. Drug Metab Dispos 1999; 27(7): 770–5
Madani S, Barilla D, Cramer J, et al. Effect of terbinafine on the pharmacokinetics and pharmacodynamics of desipramine in healthy volunteers identified as cytochrome P450 2D6 (CYP2D6) extensive metabolizers. J Clin Pharmacol 2002; 42(11): 1211–8
FDA Public Health Advisory. The safety of Sporanox® capsules and Lamisil® tablets for the treatment of onychomycosis [online]. Available from URL: http://www.fda.gov/cder/drug/advisory/sporanox-lamisil/advisory.htm [Accessed 2005 Oct 10]
Liu L, Pang KS. The roles of transporters and enzymes in hepatic drug processing. Drug Met Disp 2005; 33(1): 1–9
Venkatakrishnan K, von Moltke LL, Greenblatt DJ. Effects of the antifungal agents on oxidative drug metabolism. Clin Pharmacokinet 2000; 38(2): 111–80
Lamba JK, Lin YS, Thummel K, et al. Common allelic variants of cytochrome P4503A4 and their prevalence in different populations. Pharmacogenet 2002; 12: 121–32
Burk DL, Berghuis AM. Protein kinase inhibitors and antibiotic resistance. Pharmacol Ther 2002; 93(2-3): 283–92
Kuehl P, Zhang J, Lin Y, et al. Sequence diversity in CYP3A5 promoters and characterization of the genetic basis of polymorphic CYP3A5 expression. Nature Genet 2001; 27: 383–91
Chandra P, Brouwer KLR. The complexities of hepatic drug transport: current knowledge and emerging concepts. Pharm Res 2004; 21(5): 719–35
Faber KN, Muller M, Jansen PLM. Drug transport proteins in the liver. Adv Drug Del Rev 2003; 55: 107–24
Nishizato Y, Ieiri I, Suzuki H, et al. Polymorphisms of OATP-C (SLC21A6) and OAT3 (SLC22A8) genes: consequences for pravastatin pharmacokinetics. Clin Pharmacol Ther 2003; 73: 554–65
Seidegard J, Vorachek WR, Pero RW, et al. Hereditary differences in the expression of the human glutathione S-transferase active on trans-stillbene oxide are due to a gene deletion. Proc Natl Acad Sci U S A 1988; 85: 7293–7
Pemble S, Schroeder KR, Spencer SR, et al. Human glutathione S-transferase theta (GSTT1): cDNA cloning and the characterization of a genetic polymorphism. Biochem J 1994; 300: 271–6
Coles BF, Morel F, Rauch C, et al. Effect of polymorphism in the human glutathione S-transferase A1 promoteron hepatic GSTAi and GSTA2 expression. Pharmacogenetics 2001; 11: 663–9
Arylamine N-acetyltransferase (NAT) nomenclature [online]. Available from URL: http://www.louisville.edu/medschool/pharmacology/NAT.html [Accessed 2005 Oct 10]
Relling MV, Rubnitz JE, Rivera GK, et al. Mercaptopurine therapy intolerance and heterozygosity at the thiopurine S-methyltransferase locus. J Natl Cancer Inst 1999; 91: 2001–8
Nagata K, Yamazoe Y. Pharmacogenetics of sulfotransferase. Annu Rev Pharmacol Toxicol 2000; 40: 159–76
Freimuth RR, Eckloff B, Wieben ED, et al. Human sulfotransferase SULT1C1 pharmacogenetics: gene resequencing and functional genomic studies. Pharmacogenetics 2001; 11: 747–56
Nowell S, Sweeney C, Winters M, et al. Association between sulfotransferase 1A1 genotype and survival of breast cancer patients receiving tamoxifen therapy. J Natl Cancer Inst 2002; 94(21): 1635–40
Tukey RH, Strassburg CP. Human UDP-glucuronosyltransferases: metabolism, expression, and disease. Annu Rev Pharmacol Toxicol 2000; 40: 581–616
Desai AA, Innocenti F, Ratain MJ. UGT pharmacogenetics: implications for cancer risk and cancer therapeutics. Pharmacogenetics 2003; 13: 517–23
Riley RJ, Kenna JG. Cellular models for ADMET predictions and evaluation of drug-drug interactions. Curr Opin Drug Discov Devel 2004; 7(1): 86–99
Trowsdale J, Parham P. Defense strategies and immunity-related genes. Eur J Immunol 2004; 34: 7–17
Marsh SGE, Parham P, Barber LD. The HLA factsbook. San Diego (CA): Academic Press, 2000
Nikolich-Zugich J, Fremont DH, Miley MJ, et al. The role of MHC polymorphism in anti-microbial resistance. Microbes Infect 2004; 6: 501–12
Jeffery KJM, Bangham CRM. Do infectious diseases drive MHC diversity? Microbes Infect 2000; 2: 1335–41
Brodsky FM. Stealth, sabotage and exploitation. Immunol Rev 1999; 168: 5–11
DiMauro S, Schon E. Mechanisms of Disease: Mitochondrial respiratory-chain diseases. New Engl J Med 2003; 348(26): 2656–68
The Jackson Laboratory. JAX® mice literature [online]. Available from URL: http://jaxmice.jax.org/library/notes/498l.html [Accessed 2005 Oct 10]
The Jackson Laboratory. Mouse genome informatics [online]. Available from URL: http://www.informatics.jax.org/ [Accessed 2005 Oct 10]
Ajit C, Suvannasankha A, Zaeri N, et al. Terbinafine-associated hepatotoxicity. Am J Med Sci 2003; 325(5): 292–5
Cleary JD, Chapman SW. Altered gene expression in amphotericin B therapy [abstract 139]. 19th Annual American College of Clinical Pharmacy Meeting; 1996 Aug 4–7; Nashville (TN). 139
Cleary JD, Gordon R, Chapman SW. Phase II clinical trial of daclizumab/amphotericin B combination [abstract M-967]. 43rd Interscience Conference on Antimicrobial Agents and Chemotherapy; 2003 Sep 14–17; Chicago (IL). 447
Cleary JD, Schwartz M, Rogers PD, et al. Effects of amphotericin B and caspofungin on histamine expression. Pharmacotherapy 2003; 23(8): 966–73
Caspofungin package insert. Liberty Corner (NJ): Merck Pharmaceuticals, 2005
Cleary JD, Kelley KL, Chapman BA, et al. Differential transcriptome expression: rhodanase and cyanide clearance. 45th Interscience Conference on Antimicrobial Agents and Chemotherapy; 2005 Dec 16–19; Washington, DC
Broad Institute. Fungal genome initiative [online]. Available from URL: http://www.broad.mit.edu/annotation/fungi/fgi/ [Accessed 2005 Oct 10]
National Research Council Canada. Candida albicans Research Lab [online]. Available from URL: http://Candida.bri.nrc.ca/candida/index.cfm?.%20page%20=%20CaAnno,%20assembly%2019 [Accessed 2005 Oct 10]
Sequencing of Candida albicans at the Stanford Genome Technology Center [online]. Available from URL: http://www-sequence.stanford.edu/group/candida/ [Accessed 2005 Oct 10]
Roemer T, Jiang B, Davison J, et al. Large-scale essential gene identification in Candida albicans and applications to antifungal drug discovery. Mol Microbiol 2003; 50(1): 167–81
Bender JA, Fink GR, Lorenz MC. How C. albicans survivies in vivo: a genomic view [abstract]. Seventh American Society of Microbiology Candida and Candidiasis Conference; 2004 Mar 18–22; Austin (TX).
Giaever G, Chu AM, Ni L, et al. Functional profiling of the Saccharomyces cerevisiae genome. Nature 2002; 418(6896): 387–91
De Deken X, Raymond M. Constitutive activation of the PDR16 promoter in a Candida albicans azole-resistant clinical isolate overexpressing CDR1 and CDR2. Antimicrob Agents Chemother 2004; 48(7): 2700–3
Chen CG, Yang YL, Shih HI, et al. CaNdt80 is involved in drug resistance in Candida albicans by regulating CDR1. Antimicrob Agents Chemother 2004; 48(12): 4505–12
Clancy CJ, Cheng S, Checkley M, et al. Antibody screening identifes Candida albicans genes expressed during human oropharyngeal candidiasis that contribue to virulence at mucosal and/or deep tissue sites [abstract]. Seventh American Society of Microbiology Candida and Candidiasis Conference; 2004 Mar 18–22; Austin (TX).
DiDomenico B. Novel antifungal drugs. Curr Opin Microbiol 1999; 2: 509–15
Iraqui I, Aubert S, Cormack B, et al. EPA6P is a major adhesion responsible for biofilm formation by Candida glabrata [abstract]. Seventh American Society of Microbiology Candida and Candidiasis Conference; 2004 Mar 18–22; Austin (TX).
Bruno VM, Lopez LA, Nobile CJ, et al. Analysis of candidate azole resistance genes in C. albicans [abstract]. Seventh American Society of Microbiology Candida and Candidiasis Conference; 2004 Mar 18–22; Austin (TX).
Doedt T, Krishnamurthy S, Bockmuhl DP, et al. APSES proteins regulate morphogenesis and metabolism in Candida albicans. Mol Biol Cell 2004; 15(7): 3167–80
Yeater KM, Chandra J, Cheng G, et al. Microarray analysis of Candida glabrata gene expression during biofilm formation [abstract]. Seventh American Society of Microbiology Candida and Candidiasis Conference; 2004 Mar 18–22; Austin (TX).
Coping VM, Brown AJ, Gow NA, et al. Expression of Candida albicans genes in cells treated with antifungal agents [abstract]. Seventh American Society of Microbiology Candida and Candidiasis Conference; 2004 Mar 18–22; Austin (TX).
White TC, Holleman S, Dy F, et al. Resistance mechanisms in clinical isolates of Candida albicans. Antimicrob Agents Chemother 2002; 46(6): 1704–13
Karababa M, Coste AT, Rognon B, et al. Comparison of gene expression profiles of Candida albicans azole-resistant clinical isolates and laboratory strains exposed to drugs inducing multidrug transporters. Antimicrob Agents Chemother 2004; 48(8): 3064–79
Rogers PD, Kramer RE, Chapman SW, et al. Amphotericin B induces expression of genes encoding chemokines and cell adhesion molecules in the human monocytic cell line THP-1. J Inf Dis 2000; 180: 1259–66
Ullmann BD, Myers H, Chiranand W, et al. Inducible defense mechanism against nitric oxide in Candida albicans. Eukaryot Cell 2004; 3(3): 715–23
Cardenas ME, Cruz MC, Del Poeta M. Antifungal activities of anti-neoplastic agents: saccharomyces cerevisiae as a model system to study drug action. Clin Microbiol Rev 1999; 12: 583–611
Cunningham ML, Irwin R, Boorman G. Tox/Path team takes on differential gene expression. Environ Health Perspect 2003; 111(15): A814–5 [online]. Available from URL: http://ehp.niehs.nih.gov/txg/docs/2003/111-15/nct/nct.html [Accessed 2005 Oct 13]
Cleary JD, Sullivan D, Wilkins D, et al. Pharmacogenomics of disseminated Candidemia [abstract]. Seventh American Society of Microbiology Candida and Candidiasis Conference; 2004 Mar 18–22; Austin (TX).
Cleary JD, Chapman JW. Pharmacogenomic responses in patients with disseminated candidiasis [abstract]. International Society for Human and Animal Mycology Meeting; 2003 May 26; San Antonio (TX).
Cleary JD, Sullivan D, Fang M, et al. Anti-mycotic pharmacogenomics in candidemia [abstract M-1971]. 44th Interscience Conference on Antimicrobial Agents and Chemotherapy; 2004 Oct 30-Nov 2, Washington, DC, 440
Tang YQ, Yeaman MR, Selsted ME. Antimicrobial peptides from human platelets. Infect Immun 2002; 70(12): 6524–33
Gustafson KS, Vercellotti GM, Bendel CM, et al. Molecular mimicry in Candida albicans: role of an integrin analogue in adhesion of the yeast to human endothelium. J Clin Invest 1991; 87(6): 1896–902
Kim HS, Choi EH, Khan J, et al. Expression of genes encoding innate host defense molecules in normal human monocytes in response to Candida albicans. Infect Immun 2005; 73(6): 3714–24
Suzuki H, Kurihara Y, Takeya M, et al. A role for macrophage scavenger receptors in atherosclerosis and susceptibility to infection. Nature 1997; 386(6622): 292–6
Lilic D, Gravenor I, Robson N, et al. Deregulated production of protective cytokines in response to Candida albicans infection in patients with chronic mucocutaneous candidiasis. Infect Immun 2003; 71(10): 5690–9
Babula O, Lazdane G, Kroica J, et al. Relation between recurrent vulvovaginal candidiasis, vaginal concentrations of mannose-binding lectin, and a mannose-binding lectin gene polymorphism in Latvian women. Clin Infect Dis 2003; 37(5): 733–7
Netea MG, van Der Meer JW, Meis JF, et al. Fas-FasL interactions modulate host defense against systemic Candida albicans infection. J Infect Dis 1999; 180(5): 1648–55
Ohman SC, Jontell M, Jonsson R. Phenotypic characterization of mononuclear cells and class II antigen expression in angular cheilitis infected by Candida albicans or Staphylococcus aureus. Scand J Dent Res 1989; 97(2): 178–85
Scaringi L, Cornacchione P, Rosati E, et al. Induction and persistence in vivo of NK/LAK activity by a mannoprotein component of Candida albicans cell wall. Cell Immunol 1994; 155(2): 265–82
Mukhopadhyay K, Prasad T, Saini P, et al. Membrane sphingolipid-ergosterol interactions are important determinants of multidrug resistance in Candida albicans. Antimicrob Agents Chemother 2004; 48(5): 1778–87
Alonso R, Llopis I, Flores C, et al. Different adhesins for type IV collagen on Candida albicans: identification of a lectin-like adhesin recognizing the 7S(IV) domain. Microbiology 2000; 1147 (Pt 7): 1971–81
Netea MG, Van der Graaf C, Van der Meer JW, et al. Recognition of fungal pathogens by Toll-like receptors. Eur J Clin Microbiol Infect Dis 2004; 23(9): 672–6
Churchill GA, Airey DC, Allayee H, et al. The Collaborative Cross, a community resource for the genetic analysis of complex traits. Nat Genet 2004; 36(11): 1133–7
Hamilton PW, Thompson D, Sloan J, et al. Knowledge-guided segmentation and morphometric analysis of colorectal dysplasia. Anal Quant Cytol Histol 1995; 17: 172–82
Bartels PH, Thompson D, Bartels H, et al. Machine vision-based histometry of premalignant and malignant prostatic lesions. Pathol Res Pract 1995; 191: 935–44
Anderson NH, Hamilton PW, Bartels PH, et al. Computerized scene segmentation for the discrimination of architectural features in proliferative lesions of the breast. J Pathol 1997; 181: 374–80
Keenean S, Diamond J, McCluggage WG, et al. An automated machine vision system for the histological grading of cervical intra-epithelial neoplasia (CIN). J Pathol 2000; 192: 351–62
Diamond J, Anderson NH, Bartels PH, et al. The use of morphological characteristics and texture analysis in the identification of tissue composition in prostatic neoplasia. Hum Pathol 2004; 95: 1121–31
Acknowledgments
The authors wish to thank Drs Alice Clark, XingCong Li, Hala ElSohly, Mohammed Ilias, Erdal Bedir, Ikhlas Khan, and their colleagues at the NCNPR; Drs Mark Hamann and Jordan Zjawiony, at the Department of Pharmacognosy, for their work in isolation of the antifungal actives; Drs Li and Zjawiony for preparation of analogs; Dr Chuck Dunbar (NCNPR) for structural elucidations; Drs Melissa Jacob and Shabana Khan of the NCNPR for most of the antifungal screening. All their hard work made evaluation and illustrations of many novel compounds possible.
This paper was funded by a grant from the National Institutes of Health (NIH grant no. AI49770-01A2) entitled, “Pharmacogenomics of natural products with antifungal activity”.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Cleary, J.D., Walker, L.A. & Hawke, R.L. Antimycotic Drug Discovery in the Age of Genomics. Am J Pharmacogenomics 5, 365–386 (2005). https://doi.org/10.2165/00129785-200505060-00004
Published:
Issue Date:
DOI: https://doi.org/10.2165/00129785-200505060-00004