Abstract
Ultrasonic C scan technique has been extensively applied in nondestructive testing (NDT) in recent years. Aiming at ultrasonic C scanning original data, this paper presents a 3D reconstruction algorithm for ultrasonic C scanning image based on area iteration. Because the distance between two neighboring slices is small, the area chance between two neighboring images is also few. Therefore, we design an iterative rule on the area of the object to detect and extract the target contour of each image. After preprocessing of ultrasonic C image, we extract the contour of the object scanned by ultrasonic transducer. Then, we reconstruct the 3D ultrasonic C scanning image using marching cube algorithm. In experiments, we implement the proposed method for ultrasonic scanning data of tissue-mimicking phantom including a liver model, and we compare the proposed method with other methods. The results show that the proposed method can more accurately display the edge information of the scanned object, and improve the precision of region detection, especially suitable when the edge has the incomplete information.
You have full access to this open access chapter, Download conference paper PDF
Similar content being viewed by others
Keywords
1 Introduction
Although people discovered the fundamental physical phenomena of acoustic imaging and CT scanning in ancient times, the realization of efficient imaging device technology in the ultrasonic detection technology is hindered due to the lack of research on data acquisition and image processing in the long term [1]. With the development of science and technology, ultrasonic imaging instruments have made great progress in the past ten years. Around the field of rapid development, development of technology research on instruments, algorithms and bio-medical applications has increased exponentially. The performance of system function, image quality, imaging speed has made obvious progress. The advancement of technology has also promoted the development of practical application. The objects of ultrasound scanning have almost related to the work piece, phantom, and all the organs of the body in the medical field.
With the development of 3D imaging by CT, PET and MRI, the application of ultrasound imaging has also been extended to 3D [2]. Compared with computed tomography (CT) imaging and magnetic resonance imaging (MRI), ultrasonic imaging has the advantages of being non-invasive, non-ionizing, portable and low-cost, less harmful than radiation with imaging methods, so it’s more suitable for imaging in multi-field applications [3]. In recent years, with the development of 3D ultrasound imaging technology, it has been proved to become an increasingly important role in the practical imaging applications and reconstruction [4,5,6]. 3D ultrasound instruments are used by researchers and commercial companies as the operation of the device, and make the 3D ultrasound imaging technology integration to detection, inspection and reconstruction experiments [7,8,9,10,11]. It has been proved that ultrasound scanning equipment has been experimentally applied to the reduction of 3D ultrasound scanning phantom.
As 2D ultrasound imaging technology can only provide the scanning of tomographic image, which is unable to provide complete volume data of tissues and organs, the 3D ultrasound imaging technology has been paid attention in recent years [12]. Many 3D reconstruction algorithms focus on 2D ultrasound images. Generally, volume reconstruction methods can be classified into three categories according to their implementation [13, 14], which are Voxel based method (VBM), Pixel based method (PBM) and Function based method (FBM). After the acquisition of 3D information, the reconstruction in the usual sense is to transform every pixel in the 2D image into a 3D coordinate system, which is called Pixel based method (PBM) [15]. Another simple method of 3D reconstruction and display is based on image features.
How to improve the robustness of the reconstruction algorithm is still a challenging problem in 3D ultrasound imaging, making it more accurate and efficient to meet the needs of clinical applications. At present, there are some special difficulties in the display of 3D ultrasound images [16]. Firstly, different with CT or MRI, brightness in ultrasound image does not have the meaning of density; ultrasound image reflects the changes of acoustic impedance in the ultrasonic propagation path of the human body. Therefore, the successful method of image processing in CT or MRI cannot be simply used in the processing of ultrasound images. Then, the quality of the original 3D data will directly affect the effect of image display. The signal-to-noise ratio is low Because of the inherent speckle noise in ultrasound images, which brings difficulties to the edge detection and segmentation. Last but not least, during the acquisition of 3D ultrasound image data, it is possible to have gaps in adjacent 2D planes. If we do not use methods such as spatial interpolation, the quality of the display will be directly affected by the gap.
In order to reduce the noise and interference in process, and improve the quality of reconstruction, the image processing of ultrasonic C scan imaging should be optimized. The main purpose of the improvement is [17]: reducing speckle noise, repairing the edge and improving the efficiency of time.
For these reasons, we present a 3D reconstruction algorithm for ultrasonic C scanning image based on area iteration. An iterative rule on the area of the object is designed to detect and extract the target contour of each layer image in the proposed method. The experimental result shows that the ambient noise and the artificial disturbance are obviously weakened, and the visualization performance is strong. In the processing of irregular image and the quality of the 3D imaging focusing on ultrasonic tissue-mimicking phantom, our method is superior to the existing method.
The rest of this paper is organized as follows: Sect. 2 presents the proposed method, which includes data collecting device and the steps of our method. The proposed method is verified by real data and images in Sect. 3 and conclusions are given in Sect. 4.
2 System and Methods
2.1 Ultrasonic C Scan System
Ultrasonic C scan technology is one of the key technologies to realize qualitative, directional, quantitative and nondestructive evaluation of defects. In this work, ultrasonic C scan equipment (C-SCAN-ARS), which is produced by Allrising Technology Co. Ltd. (Beijing), is used to collect 3D data. The device is shown in Fig. 1, which is controlled by the software of the ultrasonic C scan testing system. The system made accessible to researchers in the experiment, combining with scanning software Ultrasoniclmmersed, analysis software ImageView. The ultrasonic C scan system uses a computer to control ultrasonic transducer in liver mimicking phantom mold and scans it alternately, displaying the intensity of the reflected wave in the detection distance within a specific range (phantom internal) as the brightness change continuously. The internal cross section of the mold can be drawn.
In the process of scanning phantom by ultrasonic C scan equipment, the problem of the edge loss was caused by the blurred ultrasound image and the loss of reflected wave, which should be considered and dealt with in the process of 3D imaging. However, many previous studies have focused on the image processing and reconstruction of regular objects, and it is difficult to realize the ultrasound image processing and reconstruction of irregular objects. In general, there are many factors that can affect the efficiency and quality of the ultrasound image acquisition, processing and final reconstruction. In general, the main factors reflect the following two aspects. One of the reasons is the irregular of phantom itself. There is a fluctuating, out of flatness and rough phenomenon, which has caused uneven signal intensity on the same surface. In the process of operation, the probe direction of ultrasonic C scan system remains vertical, usually forming a signal scattering phenomenon which can lead to decrease the received echo amplitude. On the other hand, image noise and interference of various types are included in scanning images, such as speckle noise, sound wave refraction, shadow interference and reverberation, which can lead to the decrease of image quality and the loss of the edge, making it difficult for the technical personnel to analyze and detect. Most of the noise interference comes from the interaction between the ultrasonic signal and the internal material.
2.2 The Proposed Method
The proposed method in this section is composed of several steps, and the flow chart is shown in Fig. 2. First, we take a series of 2D gray images from original C scanning data and preprocess each layer image, including image denoising and binarization. We then recognize all regions using 8-connected region detection method after filling empties and eliminating small holes for the binarized images. The next step is the largest area region extraction based iterative closest area algorithm. Finally, we reconstruct the 3D structure of the object using marching cube algorithm.
Firstly, we select mean filter and Gauss filter to ultrasonic C image because median filter can smooth pulse noise and protect the sharp edges of the image Gauss filter is suitable for Gauss noise. So the main methods adopted are median filter and Gauss filter method to eliminate the noise and filter.
Secondly, the image needs to be binarized to preserve the feature information of the image after image filtered. The OSTU threshold method is selected to process the smoothed image in our experiment. Here, the pixels are divided into two types: the target and the background. The result preserves the original edge features, and paves the way for the edge extraction and reconstruction subsequently.
The third step is the regions detection based 8-connected regions to each layer image. The connected area analysis has been more widely adopted in many applications of image processing and pattern recognition. This method can realize the coarse segmentation of the object identified, and the whole picture is decomposed into several connected sub-graphs to reduce the storage overhead of the system [18].
After all the connected regions are recognized, we will extract the object region in each layer image. This paper proposes the object region detect method based iterative closest area algorithm.
Let two adjacent images be \(I_{1}\) and \(I_{2}\). The multiple regions are represented by \(S=[s_{1},s_{2},\dots \,s_{i}]\) in image \(I_{1}\) and by \(W=[w_{1},w_{2},\dots \,w_{i}]\), where i and j are the number of the region detected. Figure 3 gives a diagram of two adjacent layer images.
Suppose the number of ultrasonic C scanning layers is n. That is to say, we obtain n gray images. The steps of the iterative closest area algorithm are outlined as follows.
-
1.
Selected random one binary image in the whole image set as image \(I_{1}\). If the image is the top or the bottom, the image set is traversed down or up. Let the adjacent image be \(I_{2}\).
-
2.
Estimated the areas of all the detected regions in \(I_{1}\), and marked the areas as \(s_{1}\),\(s_{2}\),\(s_{3}\) and \(s_{4}\). Likewise, the areas in \(I_{2}\) are marked by as \(w_{1}\),\(w_{2}\),\(w_{3}\),\(w_{4}\) and \(w_{5}\).
-
3.
The center of each region is computed in the areas \(s_{i}(i=1,2,3,4)\) and \(w_{j}(j=1,2,3,4,5)\). Let the center point be \(p_{i}(i=1,2,3,4)\) and \(q_{j}(j=1,2,3,4,5)\).
-
4.
As shown in Fig. 3, the maximum area \(s_{i}\) from the area set \(s_{i}\) is found in the image \(I_{1}\) and the center point corresponding to \(s_{i}\) is labeled as \(p_{1}\).
-
5.
Compared the areas \(s_{i}\) in the image \(I_{1}\) with \(t_{j}\) in the image \(I_{2}\), and estimated the area difference \({\varDelta }s_{i}\) . Because the two images are neighbor, the difference between two detection regions is very little. Therefore, we can eliminate the tiny areas, even to a few noise points.
-
6.
To determine the position of the detected region in the image \(I_{2}\), we must match the center point \(p_{1}\) of the maximum area \(s_{1}\) in the image \(I_{1}\) to that of the corresponding center point \(q_{j}(j=1,2,3,4,5)\) in the image \(I_{2}\). We first define an objective function E
$$\begin{aligned} E = ||\,{Hp_{1}-q_{j}}\, ||\,, j=(1,2,3,4,5)\; \end{aligned}$$(1)where H is a transformation matrix.
-
7.
The least squares method is used to solve the optimal solution subject min(E), and the corresponding area region in the image \(I_{2}\) can be obtained.
-
8.
Taken the previous one (or next) layer of the image processed, and repeat the above step, and so on. Iterate over the whole 3D scanning data of the object to get the result of area iteration.
Finally, we use the different methods of 3D reconstruction. The methods of 3D reconstruction are mainly divided into two groups: Surface Rendering and Direct Volume Rendering. We choose algorithms from every groups and give the results of reconstruction and performance analysis.
3 Experiment and Result
3.1 Materials and Preparation
In order to use the 3D data of liver phantom by operating the ultrasonic C scan equipment, we made a phantom including a liver model using the ultrasonic tissue-mimicking materials. The size of the phantom is 180 mm * 100 mm * 80 mm. Figure 4(a) is a liver model, and Fig. 4(b) is an ultrasound phantom to simulate the human tissue. The ultrasonic probe in the experiment was selected of the 5 MHZ phantom of water immersion ultrasonic C scan, and the phantom of liver phantom will be immersed in water for ultrasonic C scan experiment, with the scanning depth according to the maximum value of reflected wave, about 450 mm.
3.2 Two-Dimensional Image Pre-processing
The first in this experiment was acquired the one-dimensional data by ultrasonic C scan, instead of selecting the image. The scanning range in the XOY plane is 190 mm * 140 mm. The scanning step is 1 mm and the sampling rate is up to 1000 MHZ.
This experiment utilized median filtering and Gauss filtering to smooth the original image (as shown in Fig. 5(a)). First, we used the median filter of a 3 * 3 template to smooth pulse noise. Next, the image is denoised by the Gauss filter, which the variance and the mean are 3 and 2, to suppress Gauss noise. The results of the smoothing process are shown in the Fig. 5(b) and (c). Compared with the original image, the number of noise points in the results is significantly reduced.
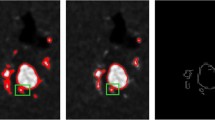
In image binarization, OSTU threshold method is selected to process the smoothed image. The binarization image is shown in the Fig. 5(d). It can be seen that the result preserves the original edge features but a few small area regions are also detected. According to the characteristics of ultrasound images, the OSTU global threshold method is used to pre-segment for the image, then the 8-connected region detect algorithm is again employed to realize twice segmentation. The experimental result of single image shows that the improved algorithm has the improvement on the level of completeness in the region detection, but also has paid attention to the speed problem in detail. The algorithm effectively measures the number of pixels in each connected domain and eliminates the noise and interference, which is not affected by the shape of the region and improve the efficiency of edge detection.
The next step is to realize the largest area detection of the scanned object through the area iteration method. We presented the iteration result of the 8 layers image in the middle part of the scanned object. The results are displayed in Fig. 6. In this experiment, the edge of the extracted image is very clear. The area iterative method overcomes the shortcomings of time consuming and complexity, which provided a method of rapid and accurate evaluation in ultrasound scan results.
3.3 Three-Dimensional Image Reconstruction
Prior to the reconstruction process, we use the two-dimensional data processing methods to ensure the useful data information can be retained and the useless data can be filtered out, which can improve the efficiency and the quality of reconstruction. In comparative experiments, we use the proposed method, the Gauss filtering and the edge detection to deal with two-dimensional data as a contrast . Then we choose three different methods to finish three-dimensional image reconstruction to verify the advantages of this proposed method. Through these different methods, different three-dimensional images can be reconstructed in different two-dimensional data processing methods.
Firstly, we choose the ISOSURFACE function in MATLAB for rendering 3D image of implicit function. In order to evaluate the reconstruction quality, we compared the reconstructed results of the Gauss filtering, the edge detection of canny, sobel and our improved algorithm. With using the method based on speckle detection and area iteration, the 3D reconstruction result of the experimental data is shown in the Fig. 7(a). The Gauss filtering reduction results are shown in Fig. 7(b). After the two simple and different kinds of edge extraction, the reduction results are shown as follows in Fig. 7(c) and (d).
Similarly, we choose a 3D volume reconstruction algorithm to restore the 3D image as the second validation. Before reconstruction can begin, in order to reduce the number of voxels to be processed in the reconstruction and to accelerate the speed of 3D reconstruction, three different methods should be used to preprocess the data set. With the useless voxel was reset to the background voxel, the 3D volume reconstruction can begin. The voxel is projected directly onto the plane and the number of projections of the data set should be calculated in this algorithm. With the algorithm, the reconstructed results of four smoothing methods can clearly generated in Fig. 8. We can use them to compare each performance on two-dimensional data filtering.
Lastly, we choose the Marching Cubes algorithm to reconstruct 3D images. The algorithm is an important method for surface reconstruction of medical images and the key is to compute the isosurface. The results of reconstructed images based on different 2D images processing are shown in Fig. 9.
After comparing and verifying the results from these reconstructed methods above, we can come to the conclusion from four algorithms of 2D data processing, that is, when we use our improved algorithm to reconstruct the 3D phantom of the ultrasonic C scan image, the noise interference has reduced and the edge contour is clear, with the overall reconstruction quality much better than the other three methods. The three different reconstructed methods can prove the same conclusion without being accidental.
Concluded from the three image reconstruction, the scan data noise mainly concentrated in the top area of liver phantom, so we focus on the phantom quality of the top with the comparison of different treatment methods for scanning data after the reconstruction, so as to evaluate reconstruction method. According to the 3D phantom with the original data of the top 50 images from 1 to 50 for the top phantom reconstruction, it can be proved that the data area at the top has aliasing noise seriously and original signal quality is poor. It is not ideal to the direct reduction of 3D phantom. By contrast, the methods of Gauss image filter and Canny edge extraction can weaken the noise interference to certain extents in Fig. 10(a) and (b), but they can’t restore the missing edge and improve the quality of reconstruction. Figure 10(c) is the 3D phantom of the top with the use of 3D image reconstruction method based on speckle detection and area iteration. The noise of the phantom is small, and the edge is clear and complete, with the phantom contour being true and accurate, which has optimized the results of image processing. At the same time, the computational time of this method is reasonable, because its image processing and iterative process are not complex, and there is no additional calculation and interference, so it is convenient to improve the quality and efficiency of the 3D reconstruction process.
4 Conclusion
Aiming at the irregular ultrasonic tissue-mimicking phantom of liver phantom, this paper has described the 3D reconstruction algorithm for ultrasonic C scanning image based on area iteration. The method calculated the contour area on the image with iterative technique according to the continuity of area, matching and extracting the contour area closest, which goes through the whole phantom data in turn and extract the entire contour, finally completing 3D reconstruction. The experimental results further verified the proposed method.
References
Sharmila, K., Sangeetha Lakshmi, G., Siva, S.A.: High-speed medical imaging in 3D ultrasound computer tomography. IRJET 2(7), 346–350 (2015)
Prager, R.W., Ijaz, U.Z., Gee, A.H., Treece, G.M.: Three-dimensional ultrasound imaging. Proc. Inst. Mech. Eng. Part H J. Eng. Med. 224(2), 193–223 (2010)
Jin, C.-Z., Nam, K.-H., Paeng, D.-G.: The spatio-temporal variation of rat carotid artery bifurcation by ultrasound imaging. In: Ultrasonics Symposium, pp. 1900–1903 (2014)
Riviere, C.N., Thakral, A., Iordachita, I.I., Mitroi, G., Stoianovici, D.: Predicting respiratory motion for active canceling during percutaneous needle insertion. In: Proceedings of the IEEE International Conference in Medicine and Biology Society, vol. 4, pp. 3477–3480 (2001)
Abolhassani, N., Patel, R.V.: Deflection of a flexible needle during insertion into soft tissue. In: Proceedings of the IEEE International Conference on Engineering in Medicine and Biology Society, vol. 1, no. 3, pp. 3858–3861 (2006)
Majewicz, A., Marra, S.P., van Vledder, M.G., Lin, M., Choti, M., Song, D.Y., et al.: Behavior of tip-steerable needles in ex vivo and in vivo tissue. IEEE Trans. Biomed. Eng. 59(10), 2705–2715 (2012)
Abayazid, M., op den Buijs, J., de Korte, C.L., Misra, S.: Effect of skin thickness on target motion during needle insertion into soft-tissue phantoms. In: IEEE Ras and Embs International Conference on Biomedical Robotics and Biomechatronics, vol. 9, no. 4, pp. 755–760 (2012)
Hong, J., Dohi, T., Hashizume, M., Konishi, K., Hata, N.: An ultrasound-driven needle-insertion robot for percutaneous cholecystostomy. Phys. Med. Biol. 49(3), 441–455 (2014)
Delgorge, C., Courreges, F., Bassit, L.A., Novales, C., Rosenberger, C., Smith-Guerin, N., et al.: A tele-operated mobile ultrasound scanner using a light-weight robot. IEEE Trans. Inf. Technol. Biomed. 9(1), 50–58 (2005)
Onogi, S., Yoshida, T., Sugano, Y., Mochizuki, T., Masuda, K.: Robotic ultrasound guidance by B-scan plane positioning control. Procedia CIRP 5(3), 100–103 (2013)
Kim, C., Chang, D., Petrisor, D., Chirikjian, G., Han, M., Stoianovici, D.: Ultrasound probe and needle-guide calibration for robotic ultrasound scanning and needle targeting. IEEE Trans. Biomed. Eng. 60(6), 1728–1734 (2013)
Pierrot, F., Dombre, E., Dégoulange, E., Urbain, L., et al.: Hippocrate: a safe robot arm for medical applications with force feedback. Med. Image Anal. 3(3), 285–300 (1999)
Solberg, O.V., Lindseth, F., Torp, H., Blake, R.E., Nagelhus Hernes, T.A.: Freehand 3D ultrasound reconstruction algorithms-a review. Ultrasound Med. Biol. 33(7), 991–1009 (2007)
Barry, C.D., Gee, A.H., Berman, L.: Three-dimensional freehand ultrasound: image reconstruction and volume analysis. Ultrasound Med. Biol. 23(8), 1209–1224 (1997)
Jensen, J.A., Nikolov, S.I., Gammelmark, K.L., Pedersen, M.H.: Synthetic aperture ultrasound imaging. Ultrasonics 1(8), 5–15 (2006)
Ungi, T., Abolmaesumi, P., Jalal, R., Welch, M., Ayukawa, I., Nagpal, S., Lasso, A., Jaeger, M., Borschneck, P.D., Fichtinger, G., Mousavi, P.: Spinal needle navigation by tracked ultrasound snapshots. IEEE Trans. Biomed. Eng. 59(10), 2766–2772 (2012)
Moon, H., Geonhwan, J., Park, S., Shin, H.: 3D freehand ultrasound reconstruction using a piecewise smooth Markov random field. Comput. Vis. Image Underst. 151, 101–113 (2016)
Suzuki, K., Horiba, I., Sugie, N.: Fast connected-component labeling based on sequential local operations in the course of forward raster scans. In: International Conference on Pattern Recognition, vol. 2, no. 2, pp. 434–437 (2000)
Acknowledgments
This work was supported in part by the National Natural Science Foundation of China under grant Nos. 61672084 and 61473025.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer Nature Singapore Pte Ltd.
About this paper
Cite this paper
Chen, B., Zhu, H., Zhou, J., Yang, P., He, L. (2017). Three-Dimensional Reconstruction of Ultrasound Images Based on Area Iteration. In: Yang, J., et al. Computer Vision. CCCV 2017. Communications in Computer and Information Science, vol 771. Springer, Singapore. https://doi.org/10.1007/978-981-10-7299-4_18
Download citation
DOI: https://doi.org/10.1007/978-981-10-7299-4_18
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-10-7298-7
Online ISBN: 978-981-10-7299-4
eBook Packages: Computer ScienceComputer Science (R0)