Abstract
Recent studies have identified patterns of genetic organization during schooling in the reproductive period of several Neotropical freshwater migratory fish. However, population segregation during non-reproductive periods is still unknown for most species. In this study, we investigated the genetic structure of populations of Salminus brasiliensis, a high-value large migratory freshwater fish, sampled during the non-reproductive season. We analysed 89 adults from Uruguay River Basin (Brazil) collected during the two consecutive non-reproductive periods, and assessed the genetic diversity levels using eleven microsatellite loci. Our results showed that populations are genetically structured, suggesting these fish can remain grouped likely due to a cooperative behaviour, not related to reproduction, in a typical shoaling behaviour. Besides, we found high genetic diversity for S. brasiliensis from the Turvo State Park area, highlighting the importance of this conservation unit as a relevant area for maintaining the genetic variability of S. brasiliensis in the Uruguay River.



Similar content being viewed by others
References
Aljanabi, S. M. & I. Martinez, 1997. Universal and rapid salt-extraction of high quality genomic DNA for PCR-based techniques. Nucleic acids research 25: 4692–4693.
Batista, J. S. & J. A. Alves-Gomes, 2006. Phylogeography of Brachyplatystoma rousseauxii (Siluriformes–Pimelodidae) in the Amazon Basin offers preliminary evidence for the first case of “homing” for an Amazonian migratory catfish. Genetics and Molecular Research 5: 723–740.
Braga-Silva, A. & P. M. Galetti, 2016. Evidence of isolation by time in freshwater migratory fish Prochilodus costatus (Characiformes, Prochilodontidae). Hydrobiologia 765: 159–167.
Carolsfeld, Y. & B. Harvey, 2003. Introduction: fishes of the floods. In Carolsfeld, J., B. Harvey, A. Baer & C. Ross (eds), Migratory fishes of South America: biology, fisheries and conservation status. International Development Research Centre and the World Bank, Victoria.
Danzmann, R. G., M. M. Ferguson & D. L. G. Noakes, 1993. The genetic basis of fish behaviour. In Pitcher, T. J. (ed.), Behaviour of Teleost Fishes. Chapman & Hall, London.
Delcourt, J. & P. Poncin, 2012. Shoals and schools: back to the heuristic definitions and quantitative references. Reviews in Fish Biology and Fisheries 22(3): 595–619.
Earl, D. & B. VonHoldt, 2012. STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conservation Genetics Resources 4: 359–361.
Evanno, G., S. Regnaut & J. Goudet, 2005. Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Molecular Ecology 14: 2611–2620.
Excoffier, L. & H. E. L. Lischer, 2010. Arlequin suite v. 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Molecular Ecology Resources 10: 564–567.
Falush, D. D., M. M. Stephens & J. K. Pritchard, 2003. Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics 164: 1567–1587.
Godoy, M. P., 1975. Peixes do Brasil, subordem Characoidei: bacia do rio Mogí Guassú. Editora Franciscana, Pirassununga.
Goudet, J., 2001. FSTAT, a program to estimate and test gene diversities and fixation indices (v. 2.9. 3).
Goulding, M., 1980. The Fishes and the Forest: Explorations in Amazonian Natural History. University of California Press, Berkeley.
Hatanaka, T. & P. M. Galetti Jr., 2003. RAPD markers indicate the occurrence of structured populations in a migratory freshwater fish species. Genetics and Molecular Biology 26: 19–25.
Jombart, T., 2008. adegenet: a R package for the multivariate analysis of genetic markers. Bioinformatics 24(11): 1403–1405.
Jombart, T., S. Devillard, A. B. Dufour & D. Pontier, 2008. Revealing cryptic spatial patterns in genetic variability by a new multivariate method. Heredity 101(1): 92–103.
Jost, L., 2008. GST and its relatives do not measure differentiation. Molelucar Ecology 17: 4015–4026.
Jueterbock, A., P. Kraemer, G. Gerlach, J. Deppermann & M. A. Jueterbock, 2012. Package “DEMEtics”. Molecular Ecology 19: 3845–3852.
Leberg, P. L., 2002. Estimating allelic richness: effects of sample size and bottlenecks. Molecular Ecology 11: 2445–2449.
Lowe-McConnell, R. H., 1967. Ecological Studies in Tropical Fish Communities. Cambridge University Press, Cambridge.
Lucas, M. C. & E. Baras, 2001. Migration of Freshwater Fishes. Blackwell Science, Oxford.
Myers, G. S., 1949. Usage of anadromous, catadromous and allied terms for migratory fishes. Copeia 1949: 89–97.
Peakall, R. & P. E. Smouse, 2006. Genalex 6: genetic analysis in Excel. Population genetic software for teaching and research. Molecular Ecology Notes 6: 288–295.
Pereira, L. H. G., F. Foresti & C. Oliveira, 2009. Genetic structure of the migratory catfish Pseudoplatystoma corruscans (Siluriformes: Pimelodidae) suggests homing behaviour. Ecology of Freshwater Fish 18: 215–225.
Pitcher, T. J., 1983. Heuristic definitions of fish shoaling behaviour. Animal Behavior 31: 611–613.
Pitcher, T. J., 1986. Functions of shoaling behaviour in teleosts. In Pitcher, T. J. (ed.), The Behaviour of Teleost Fishes. Springer, Boston.
Pitcher, T. J. & J. K. Parrish, 1993. Functions of shoaling behaviour in teleosts. In Pitcher, T. J. (ed.), Behaviour of Teleost Fishes. Chapman & Hall, London.
Pritchard, J. K., M. Stephens & P. Donnelly, 2000. Inference of population structure using multilocus genotype data. Genetics 155: 945–959.
R Development Core Team, R., 2017. R: A Language and Environment for Statistical Computing. Vienna, R Foundation for Statistical Computing.
Raymond, M. & F. Rousset, 1995. GENEPOP (version 1.2): population genetics software for exact tests and ecumenicism. Journal of heredity 86: 248–249.
Reynalte-Tataje, D. A., A. P. Nuñer, M. C. Nunes, V. Garcia, C. A. Lopes & E. Zaniboni-Filho, 2012. Spawning of migratory fish species between two reservoirs of the upper Uruguay River, Brazil. Neotropical Ichthyology 10(4): 829–835.
Ribeiro, M. B. L. B., 1983. As migrações dos jaraquis (Pisces, Prochilodontidae) no rio Negro, Amazonas, Brasil. Masters Dissertation, INPA/FUA. Manaus, 192 p.
Ribolli, J., D. J. Hoeinghaus, J. A. Johnson, E. Zaniboni-Filho, P. D. de Freitas & P. M. Galetti, 2017. Isolation-by-time population structure in potamodromous Dourado Salminus brasiliensis in southern Brazil. Conservation Genetics 18: 67–76.
Rice, W. R., 1989. Analyzing tables of statistical tests. Evolution 43: 223–225.
Rueda, E. C., P. Carriquiriborde, A. M. Monzón, G. M. Somoza & G. Ortí, 2013. Seasonal variation in genetic population structure of sábalo (Prochilodus lineatus) in the Lower Uruguay River. Genetica 141: 401–407.
Ruschel, A. R., R. O. Nodari & B. M. Moerschbacher, 2007. Woody plant species richness in the Turvo State park, a large remnant of deciduous Atlantic forest, Brazil. Biodiversity Conservation 16: 1699–1714.
Sanches, A. & P. M. Galetti Jr., 2007. Genetic evidence of population structuring in the neotropical freshwater fish Brycon hilarii (Valenciennes, 1850). Brazilian Journal of Biology 67: 889–895.
Schork, G., S. Hermes-Silva, L. F. Beux, E. Zaniboni-Filho & A. P. O. Nuñer, 2012. Diagnóstico da pesca artesanal na usina hidrelétrica de machadinho, alto Rio Uruguai – Brasil. Boletim do Instituto de Pesca 38: 97–108.
Schork, G., S. Hermes-Silva & E. Zaniboni-Filho, 2013. Analysis of fishing activity in the Itá reservoir, Upper Uruguay River, in the period 2004-2009. Brazilian Journal of Biology 73: 559–571.
Schuelke, M., 2000. An economic method for the fluorescent labeling of PCR fragments. Nature Biotechnology 18(2): 233–234.
Slatkin, M., 1995. A measure of population subdivision based on microsatellite allele frequencies. Genetics 139: 457–462.
Van Oosterhout, C., W. F. Hutchinson, D. P. M. Wills & P. Shipley, 2004. MICRO-CHECKER: software for identifying and correcting genotyping errors in microsatellite data. Molecular Ecology Notes 4: 535–538.
Weir, B. S. & C. C. Cockerham, 1984. Estimating F-statistics for the analysis of population structure. Evolution 38: 1358–1370.
Zaniboni-Filho, E., 1985. Biologia da reprodução do matrinxã, Bricon cephalus (Günther, 1869) (Teleostei: Characidae). Masters dissertation, INPA/FUA. Manaus, Brazil, 134 p.
Zaniboni-Filho, E. & U. H. Schulz, 2003. Migratory fishes of the Uruguay River. In Carolsfeld, J., B. Harvey, A. Baer & C. Ross (eds), Migratory Fishes of South America: Biology, Fisheries and Conservation Status. International Development Research Centre and the World Bank, Victoria.
Ziober, S. R., D. A. Reynalte-Tataje & E. Zaniboni-Filho, 2015. The importance of a conservation unit in a subtropical basin for fish spawning and growth. Environmental Biology of Fishes 98: 725–737.
Acknowledgements
JR acknowledges the financial support provided by Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES) and Programa de Doutorado Sanduíche no Exterior (PDSE) (process1592/81-2). PMGJ thanks Fundação de Amparo à Pesquisa do Estado de São Paulo (FAPESP, Grant 2010/52315-7), Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq, 308385/2014-4) and Sistema Nacional de Pesquisa em Biodiversidade (SISBIOTA-Brazil, MCTI/CNPq 563299/2010-0). EZF thanks Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq, Grant 302860/2014-2). We are grateful to Laboratório de Biologia e Cultivo de Peixes de Água Doce (LAPAD) of Universidade Federal de Santa Catarina (UFSC), Pedro Iaczinki and local fishermen for help with fish collections. The authors thank the two anonymous reviewers for valuable contributions improving the manuscript. Research was conducted under Animal Care Protocol PP00788.
Author information
Authors and Affiliations
Corresponding author
Additional information
Handling editor: Christian Sturmbauer
Electronic supplementary material
Below is the link to the electronic supplementary material.
10750_2018_3550_MOESM2_ESM.tif
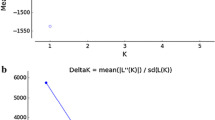
Fig. S2 Plots displays mean log-likelihood values (LnP (D)) and Evanno´s Delta K generated in STRUCTURE HARVESTER based in 89 Salminus brasiliensis from Uruguay River Basin. STRUCTURE analysis was performed with six independent runs for K = 1-10 were performed at 500,000 Markov Chain Monte Carlo (MCMC) repetitions after a burn-in period of 300,000 iterations. Delta K results indicated that samples comprise two populations (K = 2) (TIFF 17982 kb)
10750_2018_3550_MOESM3_ESM.tif
Fig. S3 Discriminant Analysis of Principal Components (DAPC). (a) Density of individual scores on the first discriminant function, with groups represented in red (Pop-2011) and blue (Pop-2011). (b) Membership probabilities (in bar plots), represent individuals in different clusters. (TIFF 119 kb)
Rights and permissions
About this article
Cite this article
Ribolli, J., Zaniboni-Filho, E., Freitas, P.D. et al. Genetic evidences of non-reproductive shoaling in the freshwater fish Salminus brasiliensis. Hydrobiologia 815, 65–72 (2018). https://doi.org/10.1007/s10750-018-3550-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10750-018-3550-y