Abstract
Cancer progression is a global burden. The incidence and mortality now reach 30 million deaths per year. Several pathways of cancer are under investigation for the discovery of effective therapeutics. The present study highlights the structural details of the ubiquitin protein ‘Ubiquitin-conjugating enzyme E2D4’ (UBE2D4) for the novel lead structure identification in cancer drug discovery process. The evaluation of 3D structure of UBE2D4 was carried out using homology modelling techniques. The optimized structure was validated by standard computational protocols. The active site region of the UBE2D4 was identified using computational tools like CASTp, Q-site Finder and SiteMap. The hydrophobic pocket which is responsible for binding with its natural receptor ubiquitin ligase CHIP (C-terminal of Hsp 70 interacting protein) was identified through protein-protein docking study. Corroborating the results obtained from active site prediction tools and protein-protein docking study, the domain of UBE2D4 which is responsible for cancer cell progression is sorted out for further docking study. Virtual screening with large structural database like CB_Div Set and Asinex BioDesign small molecular structural database was carried out. The obtained new ligand molecules that have shown affinity towards UBE2D4 were considered for ADME prediction studies. The identified new ligand molecules with acceptable parameters of docking, ADME are considered as potent UBE2D4 enzyme inhibitors for cancer therapy.















Similar content being viewed by others
References
Engblom C, Pfirschke C, Pittet JM (2016) The role of myeloid cells in cancer therapies. Nat Rev Cancer 16:447–462
Cheng M, Chen Y, Xiao W, Sun R, Tian Z (2013) NK cell-based immunotherapy for malignant diseases. Cell Mol Immunol 10:230–252
Ciechnover A (2015) The unravelling of the ubiquitin system. Nat Rev Mol Cell Biol 16:322–324
Nalepa G, Rofle M, Harper WJ (2006) Drug discovery in the ubiquitin-proteasome system. Nat Rev Drug Discov 5:596–613
Ravid T, Hochstrasser M (2008) Diversity of degradation signals in the ubiquitin-proteasome system. Nat Rev Mol Cell Biol 9:679–689
Sarita Rajender P, Ramasree D, Bhargavi K, Vasavi M, Uma V (2010) Selective inhibition of proteins regulating CDK/cyclin complexes: strategy against cancer—a review. J Recept Signal Transduct Res 30:206–213
Ciechanover A (2005) Proteolysis: from the lysosome to ubiquitin and the proteasome. Nat Rev Mol Cell Biol 6:79–87
Muratani M, Tansey PT (2003) How the ubiquitin-proteasome system controls transcription. Nat Rev Mol Cell Biol 4:192–201
Bedford L, Lowe J, Dick LR, Mayer RJ, Brownell JE (2011) Ubiquitin-like protein conjugation and the ubiquitin-proteasome system as drug targets. Nat Rev Drug Discov 10:29–46
Hu R, Hochstrasser M (2016) Recent progress in ubiquitin and ubiquitin-like protein (Ubl) signalling. Cell Res 26:389–390
Mani A, Gelmann EP (2005) The ubiquitin-proteasome pathway and its role in cancer. J Clin Oncol 23:4776–4789
Ye Y, Rape M (2009) Building ubiquitin: E2 enzymes at work. Nat Rev Mol Cell Biol 10:755–764
Berndsen CE, Wolberger C (2014) New insights into ubiquitin E3 ligase mechanism. Nat Struct Mol Biol 21:301–307
Severe N, Dieudonne FX, Marie PJ (2013) E3 ubiquitin ligase-mediated regulation of bone formation and tumorigenesis. Cell Death Disease 4:1–10
Xu Z, Kohli E, Devlin IK, Bold M, Nix CJ, Misra S (2008) Interactions between the quality control ubiquitin ligase CHIP and ubiquitin conjugating enzymes. BMC Struct Biol 8:1–13
Jiang J, Ballinger CA, Wu Y, Dai Q, Cyr DM, Hohfeld J, Patterson C (2001) CHIP is a U-box-dependent E3 ubiquitin ligase identification of Hsc70 as a target for ubiquitinylation. J Biol Chem 276:42938–42944
Kajiro M, Hirota R, Nakajima Y, Kawanowa K, So-ma K, Ito I, Yamaguchi Y, Ohie S, Kobayashi Y, Seino Y, Kawano M, Kawabe Y, Takei H, Hayashi S, Kurosumi M, Murayama A, Kimura K, Yanagisawa J (2009) The ubiquitin ligase CHIP acts as an upstream regulator of oncogenic pathways. Nat Cell Biol 11:312–319
Cavasotto NC, Phatak SS (2009) Homology modeling in drug discovery: current trends and applications. Drug Discov Today 14:676–683
Hillisch A, Pineda FL, Hilgenfeld R (2004) Utility of homology models in the drug discovery process. Drug Discov Today 9:659–669
Alpi E, Griss J, da Silva AW, Bely B, Antunes R, Zellner H, Rios D, O’Donovan C, Vizcaino JA, Martin MJ (2015) Analysis of the tryptic search space in UniProt databases. Proteomics 15:48–57
Gasteiger E, Gattiker A, Hoogland C, Ivanyi I, Appel RD, Bairoch A (2003) ExPASy: the proteomics server for in-depth protein knowledge and analysis. Nucleic Acids Res 31:3784–3788
McGinnis S, Madden TL (2004) BLAST: at the core of a powerful and diverse set of sequence analysis tools. Nucleic Acids Res 32:20–25
Ye J, McGinnis S, Madden TL (2006) BLAST: improvements for better sequences analysis. Nucleic Acids Res 34:6–9
Christian C, Jonathan DB, Geoffrey JB (2008) The Jpred 3 secondary structure prediction server. Nucleic Acids Res 36:197–201
Contreras-Moreira B, Bates PA (2002) Domain fishing: a first step in protein comparative modeling. Bioinformatics 18:1141–1142
Thompson JD, Higgins DG, Gibson TJ (1994) Clustal W: improving the sensitivity of sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Gonnet GH, Cohen MA, Benner SA (1992) Exhaustive matching of the entire protein sequence database. Science 256:1443–1445
Webb B, Sali A (2014) Comparative protein structure modeling using MODELLER. Curr Protoc Bioinformatics 47:5.6.1–5.6.32
Jacobson M, Sali A (2004) Comparative protein structure modeling and its applications to drug discovery. Annu Rep Med Chem 39:259–276
Sali A, Blundell TL (1993) Comparative modeling by satisfaction of spatial restraints. J Mol Biol 234:779–815
Banks JL, Beard HS, Cao Y, Cho AE, Damm W, Farid R, Felts AK, Halgren TA, Mainz DT, Maple JR, Murphy R, Philipp DM, Repasky MP, Zhang LY, Berne BJ, Friesner RA, Gallicchio E, Levy RM (2005) Integrated modeling program, applied chemical theory (IMPACT). J Comput Chem 26:1752–1780
Shivakumar D, Williams J, Wu Y, Damm W, Shelley J, Sherman W (2010) Prediction of absolute solvation free energies using molecular dynamics free energy perturbation and the OPLS force field. J Chem Theory Comput 6:1509–1519
Jorgensen WL, Tirado-Rives J (1996) The OPLS (optimized potentials for liquid simulations) potential functions for proteins, energy minimizations for crystals of cyclic peptides and crambin. J Am Chem Soc 110:1657–1666
Laskowski RA, MacArthur MW, Moss DS, Thornton JM (1993) PROCHEK: a program to check the stereo chemical quality of protein structures. J Appl Crystallogr 26:283–291
Zhou AQ, O’Hern CS, Regan L (2011) Revisiting the Ramachandran plot from a new angle. Prot Sci 20:1166–1171
Wiederstein M, Sippl MJ (2007) ProSA-web: interactive web service for the recognition of errors in three-dimensional structures of proteins. Nucleic Acids Res 35:407–441
Sippl MJ (1993) Recognition of errors in three-dimensional structures of proteins. Proteins 17:355–362
Kalman M, Ben-Tal N (2010) Quality assessment of protein model-structures using evolutionary conservation. Bioinformatics 26:1299–1307
Luthy R, Bowie JU, Eisenberg D (1992) Assessment of protein models with three dimensional profiles. Nature 356:83–85
Morris AL, MacArthur MW, Hutchinson EG, Thornton JM (1992) Stereochemical quality of protein structure coordinates. Proteins 12:345–364
Jiang F, Han W, YD W (2010) Influence of side chain conformations on local conformational features of amino acids implication for force field development. J Phys Chem 114:5840–5850
Sippl MJ (1995) Knowledge-based potentials for proteins. Curr Opin Struct Biol 5:229–235
Bowie JU, Luthy R, Eisenberg D (1991) A method to identify protein sequences that fold into a known three-dimensional structure. Science 253:164–170
Guex N, Peitsch MC, Schwede T (2009) Automated comparative protein structure modeling with SWISS-MODEL and Swiss-PdbViewer: a historical perspective. Electrophoresis 30:162–173
Dundas J, Ouyang Z, Seng TJ, Binkowski A, Trupaz Y, Liang J (2006) CASTp: computed atlas of surface topography of proteins with structural and topographical mapping of functionally annotated residues. Nucleic Acids Res 34:116–118
Laurie AT, Jackson RM (2005) Q-site finder: an energy-based method for the prediction of protein-ligand sites. Bioinformatics 21:1908–1916
Halgren T (2009) Identifying and characterizing binding sites and assessing druggability. J Chem Inf Mod 49:377–389
Schneidman-Duhovny D, Inbar Y, Nussinov R, Wolfson HJ (2005) PatchDock and SymmDock: servers for rigid and symmetric docking. Nucleic Acids Res 33:363–367
Binkowski TA, Naghibzadeh S, Liang J (2003) CASTp: computed atlas of surface topography of proteins. Nucleic Acids Res 31:3352–3355
Liang J, Edelsbrunner H, Woodward C (1998) Anatomy of protein pockets and cavities: measurement of binding site geometry and implications for ligand design. Prot Sci 7:1884–1897
Halgren TA (2007) New method for fast and accurate binding site identification and analysis. Chem Biol Drug Des 69:146–148
Reddy AS, Priyadarshini PS, Kumar PP, Pradeep HN, Sastry GN (2007) Virtual screening in drug discovery—a computational perspective. Curr Protein Pept Sci 8:329–351
Kitchen DB, Decornez H, Furr JR, Bajorath J (2004) Docking and scoring in virtual screening for drug discovery: methods and applications. Nat Rev Drug Discov 3:935–949
Girke T, Cheng LC, Raikhel N (2005) ChemMine. A compound mining database for chemical genomics. Plant Physiol 138:573–577
Chen IJ, Folopee N (2010) Drug-like bioactive structures and conformational coverage with the LigPrep/ConfGen suite: comparison to programs MOE and catalyst. J Chem Inf Model 50:822–839
Elokely MK, Doerksen JR (2013) Docking challenge: protein sampling and molecular docking performance. J Chem Inf Model 53:1934–1945
Friesner RA, Murphy RB, Repasky MP, Frye LL, Greenwood JR, Halgren TA, Sanschagrin PC, Mainz DT (2006) Extra precision glide: docking and scoring incorporating a model of hydrophobic encloser for protein-ligand complexes. J Med Chem 49:6177–6196
Durham E, Dorr B, Woetzel N, Staritzbichler R, Meiler J (2009) Solvent accessible surface area approximations for rapid and accurate protein structure prediction. J Mol Model 15:1093–1108
Lill MA, Danielson ML (2011) Computer-aided drug design platform using PyMOL. J Comput Aid Mol Des 25:13–19
Lipinski CA, Lombardo F, Dominy BW, Feeney PJ (1997) Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv Drug Deliv Rev 23:3–25
Ioakimids L, Thoukydidis L, Mirza A, Naeem S, Reynisson J (2008) Benchmarketing the reliability of QikProp. Correlation between experimental and predicted values. QSAR Comb Sci 27:445–456
Schwede T (2013) Protein modelling: what happened to the “protein structure gap”. Structure 21:1531–1540
Lee D, Redfern O, Orengo C (2007) Predicting protein function from sequence and structure. Nat Rev Mol Cell Biol 8:995–1005
Whisstock JC, Lesk AM (2003) Prediction of protein function from protein sequence and structure. Q Rev Biophys 36:307–340
Watson JD, Laskowski RA, Thornton JM (2005) Predicting protein function from sequence and structural data. Curr Opin Struct Biol 15:275–284
Dumpati R, Dulapalli R, Kondagari B, Ramatenki V, Vellanki S, Vadija R, Vuruputuri U (2016) Suppressor of cytokine signalling-3 as a drug target for type 2 diabetes mellitus: a structure-guided approach. Chemistry Select 1:2502–2514
Sasikala D, Jeyakanthan J, Srinivasan R (2016) Structural insights on identification of potential lead compounds targeting WbpP in vibrio vulnificus through structure-based approaches. J Recept Signal Transduct Res 36:515–530
Vadija R, Mustyala KK, Niambigari N, Dulapalli R, Dumpati RK, Ramatenki V, Vellanki SP, Vuruputuri U (2016) Homology modeling and virtual screening studies of FGF-7 protein—a structure-based approach to design new molecules against tumor angiogenesis. J Chem Biol 9:69–78
Yahalom R, Reshef D, Wiener A, Frankel S, Kalisman N, Lerner B, Keasar C (2011) Structure-based identification of catalytic residues. Proteins 79:1952–1963
Bartlett GJ, Porter CT, Borkakoti N, Thornton JM (2002) Analysis of catalytic residues in enzyme active sites. J Mol Biol 324:105–121
Mustyala KK, Malkhed V, Chittireddy VRR, Vuruputuri U (2016) Identification of small molecular inhibitors for efflux protein: DrrA of Mycobacterium tuberculosis. Cell Mol Bioeng 9:190–202
Singh T, Biswas D, Jayaram B (2011) AADS—an automated active site identification, docking and scoring protocol for protein targets based on physicochemical descriptors. J Chem Inf Model 51:2515–2527
Ramatenki V, Potlapally SR, Dumpati RK, Vadija R, Vuruputuri U (2015) Homology modeling and virtual screening of ubiquitin conjugation enzyme E2A for designing a novel selective antagonist against cancer. J Recept Signal Transduct Res 35:536–549
Malkhed V, Mustyala KK, Potlapally SR, Vuruputuri U (2014) Identification of novel leads applying in silico studies for mycobacterium multidrug resistant (MMR) protein. J Biomol Struct Dyn 32:1889–1906
Sliwoski G, Kothiwale S, Meiler J, Lowe EW Jr (2014) Computational methods in drug discovery. Pharmacol Rev 66:334–395
Lionta E, Spyrou G, Vassilatis DK, Cournia Z (2014) Structure-based virtual screening for drug discovery: principles, applications and recent advances. Curr Top Med Chem 14:1923–1938
Martins P, Jesus J, Santos S, Raposo LR, Roma-Rodrigues C, Baptista PV, Fernandes AR (2015) Heterocyclic anticancer compounds: recent advances and the paradigm shift towards the use of nanomedicine’s tool box. Molecules 20:16852–16891
Lipinski AC (2004) Lead- and drug-like compounds: the rule-of-five revolution. Drug Discov Today Technol 1:337–341
Congreve M, Carr R, Murray C, Jhoti H (2003) A ‘rule of three’ for fragment-based lead discovery? Drug Discov Today 8:876–877
Acknowledgements
The author VR acknowledges the Council of Scientific and Industrial Research, India, for the financial support (File No: 09/132(0821)/2012-EMR-I). The authors VR, RKD, RV, SPV and RR acknowledge the Principal and Head, Department of Chemistry, University College of Science, Osmania University, Hyderabad, for providing facilities to carry out this work.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethical standards
The authors state that no human studies and no animal studies were carried out for this article.
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Supplementary fig S1
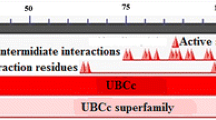
Alignment of the UBE2D4 sequence with the other E2 sequences. The alignment file was generated using CLUSTALW server tool. The UBC domain of UBE2D4 is conserved in all the E2s. The UBE2D4 conserved domain sequence starts from LYS 4 and ends with THR 142 amino acid residue. (GIF 7718 kb)
Supplementary Table S1
(DOCX 17 kb)
Supplementary Table S2
(DOCX 100 kb)
Rights and permissions
About this article
Cite this article
Ramatenki, V., Dumpati, R., Vadija, R. et al. Targeting the ubiquitin-conjugating enzyme E2D4 for cancer drug discovery–a structure-based approach. J Chem Biol 10, 51–67 (2017). https://doi.org/10.1007/s12154-016-0164-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12154-016-0164-6