Abstract
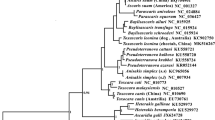
The present study was designed to determine and analyze the mt genomes of Metastrongylus salmi (M. salmi), and reveal the phylogenetic relationships of this parasite using mt DNA sequences. Results showed that the complete mt genome of M. salmi was 13722 bp containing 12 protein-coding genes (cox1-3, nad1-6, nad4L, atp6 and cytb), 22 transfer RNA genes, and 2 ribosomal RNA genes (rrnL and rrnS). The overall A+T content was 73.54% and the nucleotide composition was A (23.52%), C (6.14%), G (19.60%), T (50.02%), and N (UCAG) (0.73%). A total of 4237 amino acids are encoded from the Tibetan isolates of M. salmi mt genomes. The ATA was predicted as the most common starting codon with 41.7% (5/12 protein genes); and 11 of the 12 protein genes were found to have a TAG or TAA translation termination codon. By clustering together the phylogenetic trees of Tibetan M. salmi and Austrian M. salmi, the M. salmi isolated from Tibetan pigs was found to be highly homological with that stemmed from Austrian one. This information provides meaningful insights into the phylogenetic position of the M. salmi China isolate and represents a useful resource for selecting molecular markers for diagnosis and population studies.
Similar content being viewed by others
References
Altschul, S.F., Gish, W., Miller, W., Myers, E.W., Lipman D.J. 1990. Basic local alignment search tool. Journal of Molecular Biology, 215, 403–410
Ashburner, M., Ball, C.A., Blake, J.A., Botstein, D., Butler, H., Cherry J.M., et al. 2000. Gene Ontology: tool for the unification of biology. Nature Genetics, 25, 25–29
Benson G. 1999 Tandem repeats finder: a program to analyze DNA sequences. Nucleic Acids Research, 27, 573–580
Borgstrom, E., Lundin, S., Lundeberg J. 2011. Large scale library generation for high throughput sequencing. PLoS One, 6, e19119. DOI: 10.1371/journal.pone.0019119
Chiaromonte, F., Yap, V.B., Miller W. 2002. Scoring pairwise genomic sequence alignments. Pacific Symposium on Biocomputing, 7, 115–126
Cronn, R., Liston, A., Parks, M., Gernandt, D.S., Shen R.K. 2008. Multiplex sequencing of plant chloroplast genomes using Solexa sequencing-by-synthesis technology. Nucleic Acids Research, 36, e122
Durent, L., Mouchiroud D. 1999. Expression pattern and, surprisingly, gene length shape codon usage in Caenorhabditis, Drosophila and Arabidopsis. Proceedings of the National Academy of Sciences of the United States of America, 96, 4482–4487
García-González Á.M., Pérez-Martín, J.E., Gamito-Santos, J.A., Calero-Bernal, R., Alonso, M.A., Carrión E.M. 2013. Epidemiologic study of lung parasites (Metastrongylus spp.) in wild boar (Sus scrofa) in southwestern Spain. Journal of Wildlife Disease, 49, 157–162. DOI: 10.7589/2011-07-217
Gassó D., Rossi, L., Mentaberre, G., Casas, E., Velarde, R., Nosal, P., et al. 2014. An identification key for the five most common species of Metastrongylus. Parasitology Research, 113, 3495–3500. DOI: 10.1007/s00436-014-4001-y
Helfenbein, K.G., Brown, W.M., Boore J.L. 2001. The complete mitochondrial genome of the articulate brachiopod Terebratalia transversa. Molecular Biology and Evolution, 18, 1734–1744
Hu, M., Chilton, N.B., Gasser R.B. 2003. The mitochondrial genome of Strongyloides stercoralis (Nematoda)-idiosyncratic gene order and evolutionary implications. International Journal of Parasitology, 33, 1393–1408
Humbert, J.F., Drouet J. 1990. Enquête épidémiologique sur la métastrongylose du sanglier (Sus scrofa) en France. Gibier Faune Sauvag, 7, 67–84. (In French)
Jex, A.R., Hall, R.S., Littlewood, D.T., Gasser R.B. 2010. An integrated pipeline for next-generation sequencing and annotation of mitochondrial genomes. Nucleic Acids Research, 38, 522–533
Kanehisa M. 1997. A database for post-genome analysis. Trends in Genetics, 13, 375
Kanehisa, M., Goto, S., Hattori, M., Aoki-Kinoshita, K.F., Itoh, M., Kawashima, S., et al. 2006. From genomics to chemical genomics: new developments in KEGG. Nucleic Acids Research, 34, D354–D357
Kanehisa, M., Goto, S., Kawashima, S., Okuno, Y., Hattori M. 2004. The KEGG resource for deciphering the genome. Nucleic Acids Research, 32, D277–D280
Kurtz, S., Phillippy, A., Delcher, A.L., Smoot, M., Shumway M., Antonescu, C., Salzberg S.L. 2004. Versatile and open software for comparing large genomes. Genome Biology, 5, R12
Li, H., Handsaker, B., Wysoker, A., Fennell, T., Ruan, J., Homer, N., et al. 2009. The Sequence Alignment/Map (SAM) format and SAMtools. Bioinformatics, 25, 2078–2079
Li, K., Lan, Y.F., Luo, H.Q., Shahzad, M., Zhang, H., Wang, L., et al. 2017a. Prevalence of three Oesophagostomum spp. from Tibetan pigs analyzed by genetic markers of nad1, cox3 and ITS1. Acta Parasitologica, 62, 90–96. DOI: 10.1515/ap-2017-0010
Li, K., Luo, H.Q., Zhang, H., Lan, Y.F., Han, Z.Q., Shahzad, M., et al. 2016. First report of Metastrongylus pudendotectus by the genetic characterization of mitochondria genome of cox1 in pigs from Tibet, China. Veterinary Parasitology, 223, 91–95. DOI: 10.1016/j.vetpar.2016.04.036
Li, K., Zhang, L.H., Zhang, H., Lei, Z.X., Luo, H.Q., Mehmood, K., et al. 2017b. Epidemiology investigation and risk factors of Echinococcus granulosus in yaks (Bos grunniens), Tibetan pigs and Tibetans on the Qinghai Tibetan plateau. Acta Tropica 173, 147–152. DOI: 10.1016/j.actatropica.2017.06.019
Li, M.W., Lin, R.Q., Song, H.Q., Wu, X.Y., Zhu X.Q. 2008. The complete mitochondrial genomes for three Toxocara species of human and animal health significance. BMC Genomics, 9, 224
Li, R., Zhu, H., Ruan, J., Qian, W., Fang, X., Shi, Z., et al. 2010. De novo assembly of human genomes with massively parallel short read sequencing. Genome Research, 20, 265–272
Li X.R. (Ed.) 2011. Color atlas of animal parasitosis (2nd Edition). Beijing: China Agriculture Press
Lin, R.Q., Liu, G.H., Hu, M., Song, H.Q., Wu, X.Y., Li, M.Y., et al. 2012. Oesophagostomum dentatum and Oesophagostomum quadrispinulatum: c haracterization of the complete mitochondrial genome sequences of the two pig nodule worms. Experimental Parasitology 131, 1–7. DOI: 10.1016/j.exppara.2012.02.015
Lohse, M., Drechsel, O., Bock R. 2007. Organellar Genome DRAW (OGDRAW): a tool for the easy generation of high-quality custom graphical maps of plastid and mitochondrial genomes. Current Genetics, 52, 267–274
Magrane M. 2011. UniProt. Knowledgebase: a hub of integrated protein data. Databases-Oxford, 2011, 9. DOI: 10.1093/database/bar009
Marruchella, G., Paoletti, B., Speranza, R., Di Guardo G. 2012. Fatal bronchopneumonia in a Metastrongylus elongatus and Porcine circovirus type 2 co-infected pig. Research in Veterinary Science, 93, 310–312. DOI: 10.1016/j.rvsc.2011.05.016
Romero, H., Zavala, A., Musto H. 2000. Codon usage in Chlamydia trachomatis is the result of strand-specific mutational biases and a complex pattern of selective force. Nucleic Acids Research, 28, 2084–2090
Saha, S., Bridges, S., Magbanua, Z.V., Peterson D.G. 2008. Empirical comparison of ab initio repeat finding programs. Nucleic Acids Research, 36, 2284–2294
Singer, G.A., Hickey D.A. 2000. Nucleotide bias causes a genomewide bias in the amino acid composition of proteins. Molecular Biology and Evolution, 17, 1581–1588
Sorensen, M., Sanz, A., Gómez, J., Pamplona, R., Portero-Otín, M., Gredilla, R., Barja G. 2006. Effects of fasting on oxidative stress in rat liver mitochondria. Free Radical Research, 40, 339–347
Tannistha, N., Catherine, O., Arvind, P.S., Boddey, J., Atkins, T., Sarkar-Tyson, M., et al. 2010. A genomic survey of positive selection in Burkholderia pseudomallei provides insights into the evolution of accidental virulence. PLoS Pathogens, 6, 1–15
Tatusov, R.L., Fedorova, N.D., Jackson, J.D., Jacobs, A.R., Kiryutin B., Koonin, E.V., et al. 2003. The COG database: an updated version includes eukaryotes. BMC Bioinformatics, 4, 41
Tatusov, R.L., Koonin, E.V., Lipman D.J. 1997. A genomic perspective on protein families. Science, 278, 631–637
Wyman, S.K., Jansen, R.K., Boore J.L. 2004. Automatic annotation of organellar genomes with DOGMA. Bioinformatics, 20, 3252–3255
Zhang, N.Z., Zhou, D.H., Huang, S.Y., Wang, M., Shi, X.C., Ciren, D.B., Zhu X.Q. 2014. Seroprevalence and risk factors associated with Haemophilus parasuis infection in Tibetan pigs in Tibet. Acta Tropica, 132, 94–97. DOI: 10.1016/j.actatropica.2013.12.021
Zhao, G.H., Hua, B., Cheng, W.Y., Jia, Y.Q., Li, H.M., Yu, S.K., Liu G.H. 2013. The complete mitochondrial genomes of Oesophagostomum asperum and Oesophagostomum columbianum in small ruminants. Infection. Infection Genetics and Evolution, 19, 205–211. DOI: 10.1186/1756-3305-7-319
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Li, K., Shahzad, M., Zhang, H. et al. Characterization of the complete mitochondrial genome of Metastrongylus salmi (M. salmi) derived from Tibetan pigs in Tibet, China. Acta Parasit. 63, 280–286 (2018). https://doi.org/10.1515/ap-2018-0032
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1515/ap-2018-0032